IB Biology SL (Standard level)- 2024 – Practice Questions- All Topics
Topic 2.5 Enzymes
Topic 2 Weightage : 20%
All Questions for Topic 2.5-Enzyme & Substrate, Enzyme Catalysis, Enzyme Specificity, Enzyme Activity, Enzyme Experiments, Enzymes in Industry, Models of Action, Types of Enzymes, Lactose Intolerance
Question
Which enzyme is matched to its function?
Enzyme | Function |
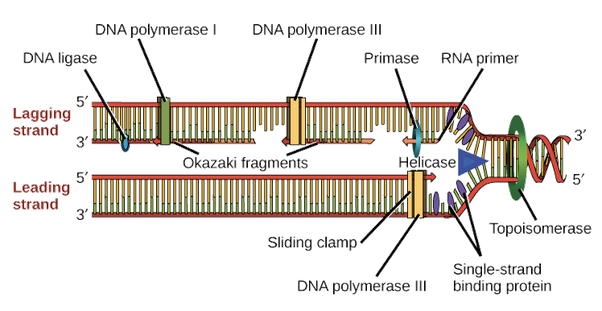
helicase | forms a DNA helix |
DNA polymerase | forms a covalent bond between DNA nucleotides |
restriction endonuclease | seals nicks in recombinant DNA |
ligase | unwinds the double helix ▶️Answer/ExplanationAns: B
DNA polymerase is an enzyme that synthesizes DNA molecules from deoxyribonucleotides, the building blocks of DNA. These enzymes are essential for DNA replication and usually work in pairs to create two identical DNA strands from a single original DNA molecule. The main function of DNA polymerase is to make DNA from nucleotides, the building blocks of DNA. |
Question
Which statement applies to enzymes?
Enzyme function depends on collisions between substrate and active sites.
One active site typically binds to a broad range of substrates.
The active site on the substrate is specific to one enzyme.
When enzymes are immobilized they stop working.
▶️Answer/Explanation
Ans: A
Enzymes are proteins that act as biological catalysts, meaning they speed up chemical reactions in living cells. Enzymes function by binding to one or more molecules called substrates, which are the reactants in the reaction. The part of the enzyme that binds the substrate is called the active site
Enzyme function depends on collisions between substrate and active site because only when the substrate collides with the active site can it form an enzyme-substrate complex. This complex is where the enzyme lowers the activation energy of the reaction and facilitates the conversion of substrate to product. The more collisions there are between substrate and active site, the faster the reaction rate.
However, not every collision between substrate and active site leads to a successful reaction. The substrate must have a shape and charge that is complementary to the active site. This is called enzyme specificity. Enzymes are very specific to their substrates and usually only catalyze one type of reaction. Some enzymes may bind to a broad range of substrates, but they usually have a preference for one or a few substrates that fit best in their active site.
The binding of substrate and active site is also influenced by environmental factors such as temperature, pH, and concentration. These factors can affect the shape and charge of both the enzyme and the substrate, and thus affect their ability to bind. For example, increasing the temperature can increase the kinetic energy of the molecules and cause more collisions between substrate and active site, but if the temperature is too high, it can also denature the enzyme and change its shape. Similarly, changing the pH can alter the charge of the amino acids in the active site and affect their attraction to the substrate.
Therefore, enzyme function depends on collisions between substrate and active site, but also on other factors that affect the binding affinity and catalytic activity
Three flasks were prepared for an analysis of the activity of amylase. At time zero, each of the substances indicated in the diagrams was added.
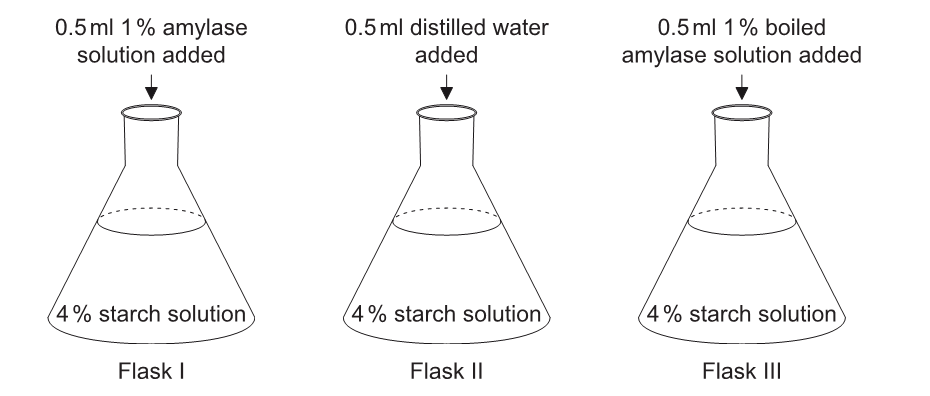
Which flask(s) could provide support for the hypothesis that heat denatures enzymes?
A. Flasks I and II after 15 minutes
B. Flasks II and III after 15 minutes
C. Flasks I and III after 15 minutes
D. Flask III at time zero and again after 15 minutes
▶️Answer/Explanation
Markscheme
C
Question
The bacterium Neisseria gonorrhoeae causes infections related to the human reproductive system. The graph shows the percentage of samples in which this bacterium showed resistance to six antibiotics over a period of ten years.
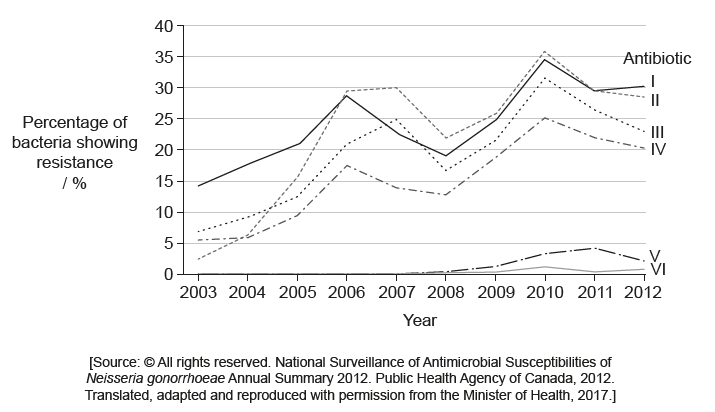
What is a possible explanation for the total percentage resistance being larger than 100% in 2010?
A. People do not take the antibiotics as prescribed.
B. More people have been sampled in that year.
C. There was an epidemic of Neisseria gonorrhoeae in that year.
D. Some bacteria are resistant to more than one antibiotic.
▶️Answer/Explanation
Markscheme
D
Question
What happens as an enzyme becomes denatured?
A. The enzyme works faster.
B. The enzyme works slower.
C. The enzyme can perform a new role.
D. The enzyme can make the reverse reaction proceed faster.
▶️Answer/Explanation
Markscheme
Ans:B
Enzymes are proteins that catalyze chemical reactions in living organisms. They have a specific three-dimensional structure that allows them to interact with other molecules and speed up chemical reactions. When an enzyme denatures, its structure changes and it can no longer interact with other molecules in the same way. This can cause the rate of reactions to slow down or even stop.
On denaturation, the shape of the active sites by the enzymes gets deformed or altered due to which the substrate does not fit into the assigned enzymes anymore. This leads to slowing down of the rate of reactions or even stopping.
Factors such as high temperatures can cause an enzyme to lose its shape (denature) and stop working. Each enzyme has an optimum pH range. Extreme high temperatures can cause an enzyme to lose its shape (denature) and stop working.
Question
The activity of amylase from two bacterial species and a fungus was measured at different pH levels and constant temperature. The results are shown in the graph.
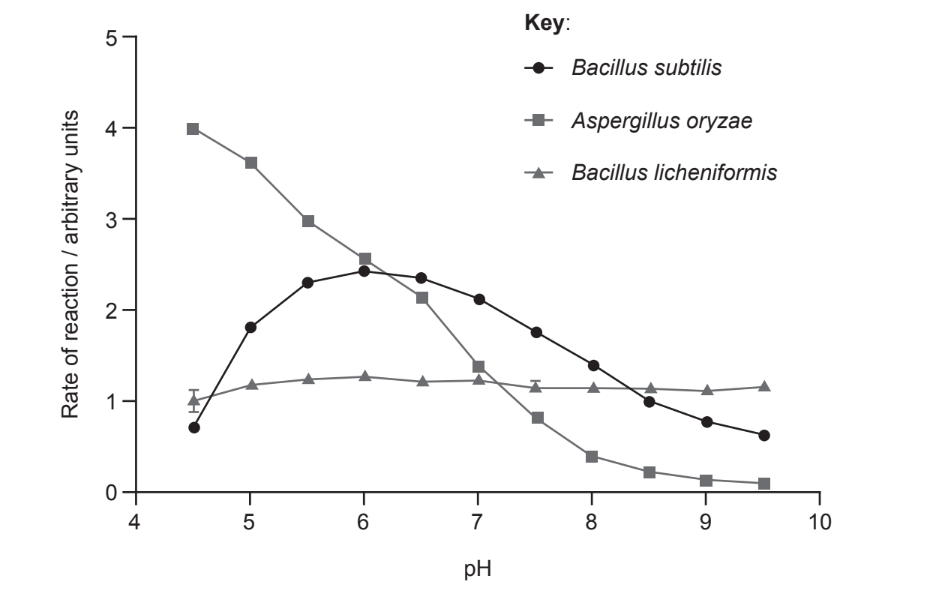
Which statement about the effect of pH on amylase can be concluded?
A. A. oryzae amylase has the highest optimum pH.
B. A change in pH affects amylase most in B. licheniformis.
C. The optimum pH is 6 in B. subtilis.
D. Amylase activity at pH 8 is the lowest in B. licheniformis.
▶️Answer/Explanation
Ans:D
Amylase is an enzyme that breaks down starch into sugars. It is produced by many organisms, including bacteria such as Bacillus licheniformis. The activity of amylase depends on several factors, such as temperature, substrate concentration, and pH. The optimal pH for amylase activity varies depending on the source of the enzyme. For B. licheniformis, the optimal pH is around 6.5, and the enzyme becomes less active at higher or lower pH values. This may be due to changes in the electrostatic interactions or the conformation of the enzyme at different pH levels. Some mutations in the gene encoding amylase can also affect the pH profile of the enzyme.