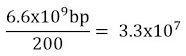Molecular basis of Inheritance : Notes and Study Materials -pdf
Notes and Study Materials
- Concepts of Molecular basis of Inheritance
- Master File Molecular basis of Inheritance
- NCERT Book chapter Molecular basis of Inheritance
- NCERT Solutions for – Molecular basis of Inheritance
- NCERT Exemplar Solutions for – Molecular basis of Inheritance
- Revision Note of Molecular basis of Inheritance
- Past Many Years Question papers and Answer of Molecular basis of Inheritance
- Mind Map of Molecular basis of Inheritance
Examples and Exercise
6. MOLECULAR BASIS OF INHERITANCE
· Nucleic acids (DNA & RNA) are the building blocks of genetic material.
· DNA is the genetic material in most of the organisms.
· RNA is the genetic material in some viruses. RNA mostly functions as messengers.
THE DNA
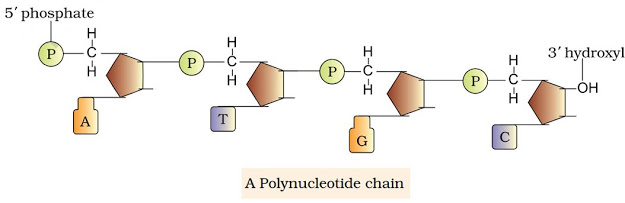
STRUCTURE OF POLYNUCLEOTIDE CHAIN
Polynucleotides are the polymer of nucleotides. DNA & RNA are polynucleotides. A nucleotide has 3 components:
1. A nitrogenous base.
2. A pentose sugar (ribose in RNA & deoxyribose in DNA).
3. A phosphate group.
Nitrogen bases are 2 types:
} Purines: It includes Adenine (A) and Guanine (G).
} Pyrimidines: It includes Cytosine (C), Thymine (T) & Uracil (U). Thymine (5-methyl Uracil) present only in DNA and Uracil only in RNA.
A nitrogenous base is linked to the OH of 1′ C pentose sugar through an N-glycosidic linkage to form nucleoside.
Nucleosides in RNA | Nucleosides in DNA |
Adenosine | Deoxyadenosine |
Guanosine | Deoxyguanosine |
Cytidine | Deoxycytidine |
Uridine | Deoxythymidine |
A phosphate group is linked to OH of 5′ C of a nucleoside through phosphoester linkage to form nucleotide (or deoxynucleotide).
In RNA, each nucleotide has an additional –OH group at 2‘ C of the ribose (2’- OH).
2 nucleotides are linked through 3’-5’ phosphodiester bond to form dinucleotide.
When more nucleotides are linked, it forms polynucleotide.
STRUCTURE OF THE DNA
} Friedrich Meischer (1869): Identified DNA and named it as ‘Nuclein’.
} James Watson & Francis Crick (1953) proposed double helix model of DNA. It was based on X-ray diffraction data produced by Maurice Wilkins & Rosalind Franklin.
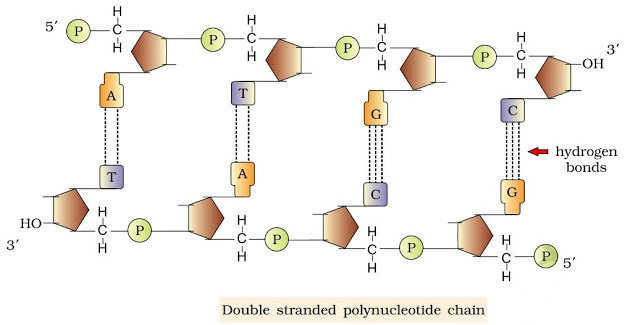
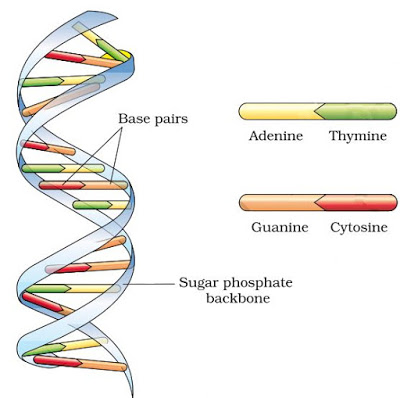
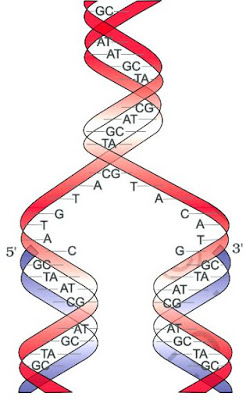
} DNA is made of 2 polynucleotide chains coiled in a right-handed fashion. Its backbone is formed of sugar & phosphates. The bases project inside.
} The 2 chains have anti-parallel polarity, i.e. one chain has the polarity 5’→3’ and the other has 3’→5’.
} The bases in 2 strands are paired through H-bonds forming base pairs (bp).
A=T (2 hydrogen bonds) C≡G (3 hydrogen bonds)
} Purine comes opposite to a pyrimidine. This generates uniform distance between the 2 strands.
} Erwin Chargaff’s rule: In DNA, the proportion of A is equal to T and the proportion of G is equal to C.
[A] + [G] = [T] + [C] or [A] + [G] / [T] + [C] =1
v Ф 174 (a bacteriophage) has 5386 nucleotides.
v Bacteriophage lambda has 48502 base pairs (bp).
v E. coli has 4.6×106 bp.
v Haploid content of human DNA is 3.3×109 bp.
Length of DNA = number of base pairs X distance between two adjacent base pairs.
Number of base pairs in human = 6.6 x 109
Hence, the length of DNA = 6.6 x109 x 0.34x 10-9
= 2.2 m
In E. coli, length of DNA =1.36 mm (1.36 x 10-3 m)
Therefore the number of base pairs
PACKAGING OF DNA HELIX
§ In prokaryotes (E.g. E. coli), the DNA is not scattered throughout the cell. DNA is negatively charged. So it is held with some positively charged proteins to form nucleoid.
§ In eukaryotes, there is a set of positively charged, basic proteins called histones.
§ Histones are rich in positively charged basic amino acid residues lysines and arginines.
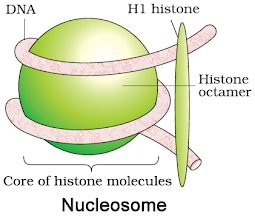
§ 8 histones form histone octamer.
§ Negatively charged DNA is wrapped around histone octamer to give nucleosome.
§ A typical nucleosome contains 200 bp.
Therefore, total number of nucleosomes in human =
§ Nucleosomes constitute the repeating unit to form chromatin. Chromatin is the thread-like stained bodies.
§ Nucleosomes in chromatin = ‘beads-on-string’.
§ Chromatin is packaged → chromatin fibres → coiled and condensed at metaphase stage → chromosomes.
§ Higher level packaging of chromatin requires non-histone chromosomal (NHC) proteins.
§ Chromatin has 2 forms:
· Euchromatin: Loosely packed and transcriptionally active region of chromatin. It stains light.
· Heterochromatin: Densely packed and inactive region of chromatin. It stains dark.
THE SEARCH FOR GENETIC MATERIAL
Frederick Griffith used mice & Streptococcus pneumoniae.
Streptococcus pneumoniae has 2 strains:
◦ Smooth (S) strain (Virulent): Has polysaccharide mucus coat. Cause pneumonia.
◦ Rough (R) strain (Non-virulent): No mucus coat. Do not cause Pneumonia.
Experiment:
- S-strain → Inject into mice → Mice die
- R-strain → Inject into mice → Mice live
- S-strain (Heat killed) → Inject into mice → Mice live
- S-strain (Heat killed) + R-strain (live) → Inject into mice → Mice die
He concluded that some ‘transforming principle’ transferred from heat-killed S-strain to R-strain. It enabled R-strain to synthesize smooth polysaccharide coat and become virulent. This must be due to the transfer of genetic material.
2. Biochemical characterization of transforming principle
– Oswald Avery, Colin MacLeod & Maclyn McCarty worked to determine the biochemical nature of ‘transforming principle’ in Griffith’s experiment.
– They purified biochemicals (proteins, DNA, RNA etc.) from heat killed S cells using suitable enzymes.
– They discovered that
- Digestion of protein and RNA (using Proteases and RNases) did not affect transformation. It means that the transforming substance was not a protein or RNA.
- Digestion of DNA with DNase inhibited transformation. It means that DNA caused transformation of R cells to S cells. It proves that DNA was the transforming principle.
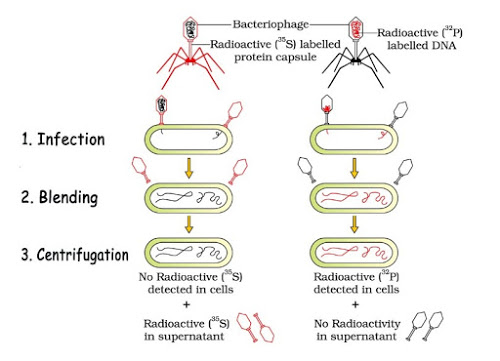
3. Hershey-Chase Experiment (Blender Experiment)-1952
- Hershey & Chase grew some bacteriophage viruses on a medium containing radioactive phosphorus (P32) and some others on medium containing radioactive sulphur (S35).
- Viruses grown in P32 got radioactive DNA because only DNA contains phosphorus. Viruses grown in S35 got radioactive protein because protein contains sulphur.
- These preparations were used separately to infect E. coli.
- After infection, the E. coli cells were gently agitated in a blender to remove the virus particles from the bacteria.
- Then the culture was centrifuged to separate lighter virus particles from heavier bacterial cells.
- Bacteria infected with viruses having radioactive DNA were radioactive. i.e., DNA had passed from the virus to bacteria. Bacteria infected with viruses having radioactive proteins were not radioactive. i.e., proteins did not enter the bacteria from the viruses. This proves that DNA is the genetic material.
PROPERTIES OF GENETIC MATERIAL (DNA v/s RNA)
A genetic material must have the following properties:
- Ability to generate its replica (Replication).
- Chemical and structural stability.
- Provide the mutations that are required for evolution.
- Ability to express as Mendelian Characters.
Reasons for stability (less reactivity) of DNA
Reasons for mutability (high reactivity) of RNA
Double stranded
Single stranded
Presence of thymine
Presence of Uracil
Absence of 2’-OH in sugar
Presence of 2’-OH in sugar
– RNA is unstable. So, RNA viruses (E.g. Q.B bacteriophage, Tobacco Mosaic Virus etc.) mutate and evolve faster.
– DNA strands are complementary. On heating, they separate. In appropriate conditions, they come together. In Griffith’s experiment, some properties of DNA of the heat killed bacteria did not destroy. It indicates the stability of DNA.
– For the storage of genetic information, DNA is better due to its stability. But for the transmission of genetic information, RNA is better.
– RNA can directly code for the protein synthesis, hence can easily express the characters. DNA is dependent on RNA for protein synthesis.
RNA WORLD
- RNA was the first genetic material.
- It acts as genetic material and catalyst.
- Essential life processes (metabolism, translation, splicing etc.) evolved around RNA.
- DNA evolved from RNA for stability.
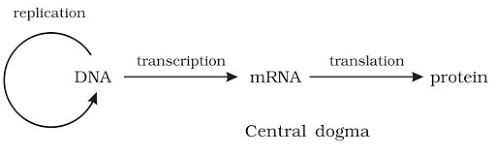
CENTRAL DOGMA OF MOLECULAR BIOLOGY
· It is proposed by Francis Crick. It states that the genetic information flows from DNA → RNA → Protein.
· In some viruses, flow of information is in reverse direction (from RNA to DNA). It is called reverse transcription.
DNA REPLICATION
· Replication is the copying of DNA from parental DNA.
· Watson & Crick proposed Semi-conservative model of replication. It suggests that the parental DNA strands act as template for the synthesis of new complementary strands. After replication, each DNA molecule would have one parental and one new strand.
· Matthew Messelson & Franklin Stahl (1958) experimentally proved Semi-conservative model.
Messelson & Stahl’s Experiment
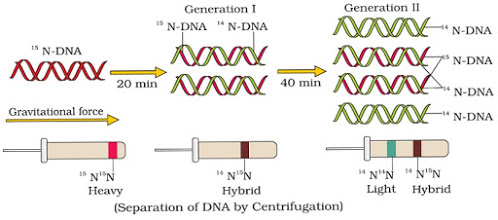
} They grew E. coli in 15NH4Cl medium (15N = heavy isotope of nitrogen) as the only nitrogen source. As a result, 15N was incorporated into newly synthesised DNA (heavy DNA or 15N DNA).
} Heavy DNA can be distinguished from normal DNA (light DNA or 14N DNA) by centrifugation in a cesium chloride (CsCl) density gradient.
} E. coli cells from 15N medium were transferred to 14NH4Cl medium. After one generation (i.e. after 20 minutes), they isolated and centrifuged the DNA. Its density was intermediate (hybrid) between 15N DNA and 14N DNA. This shows that in newly formed DNA, one strand is old (15N type) and one strand is new (14N type). This confirms semi-conservative replication.
} After II generation (i.e. after 40 minutes), there was equal amounts of hybrid DNA and light DNA.
Taylor & colleagues (1958) performed similar experiments on Vicia faba (faba beans) using radioactive thymidine to detect distribution of newly synthesized DNA in the chromosomes. It proved that the DNA in chromosomes also replicate semi-conservatively.
The Machinery and Enzymes for Replication
· DNA replication starts at a point called origin (ori).
· A unit of replication with one origin is called a replicon.
· During replication, the 2 strands unwind and separate by breaking H-bonds in presence of an enzyme, Helicase.
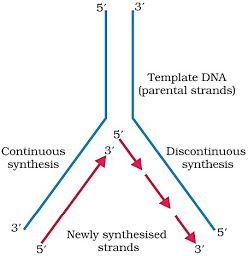
· Unwinding of the DNA molecule at a point forms a ‘Y’-shaped structure called replication fork.
 |
| Watson-Crick model for semiconservative DNA replication |
· The separated strands act as templates for the synthesis of new strands.
· DNA replicates in the 5’→3’ direction.
· Deoxyribonucleoside triphosphates (dATP, dGTP, dCTP & dTTP) act as substrate and provide energy for polymerization.
· Firstly, a small RNA primer is synthesized in presence of an enzyme, primase.
· In presence of an enzyme, DNA dependent DNA polymerase, many nucleotides join with one another to primer strand and form a polynucleotide chain (new strand).
· During replication, one strand is formed as a continuous stretch in 5’→ 3’ direction (Continuous synthesis). This strand is called leading strand.
· The other strand is formed in small stretches (Okazaki fragments) in 5’→ 3’ direction (Discontinuous synthesis).
· The Okazaki fragments are then joined together to form a new strand by an enzyme, DNA ligase. This new strand is called lagging strand.
· If a wrong base is introduced in the new strand, DNA polymerase can do proof reading.
· E. coli completes replication within 18 minutes. i.e. 2000 bp per second.
· In eukaryotes, the replication of DNA takes place at S-phase of the cell cycle. Failure in cell division after DNA replication results in polyploidy.
TRANSCRIPTION
– It is the process of copying genetic information from one strand of the DNA into RNA.
– Here, adenine pairs with uracil instead of thymine.
– The DNA- dependent RNA polymerase catalyzes the polymerization only in 5’→3’direction.
– 3’→5’ acts as template strand. RNA is built from this.
– 5’→3’ acts as coding strand. This is copied to RNA.
3’-ATGCATGCATGCATGCATGCATGC-5’ template strand.
5’-TACGTACGTACGTACGTACGTACG-3’ coding strand.
– During transcription, both strands are not copied because
◦ The code for proteins is different in both strands. This complicates the translation.
◦ If 2 RNA molecules are produced simultaneously, this would be complimentary to each other. It forms a double stranded RNA and prevents translation.
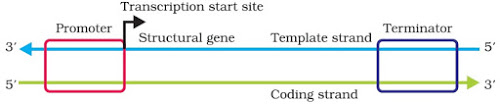
Transcription Unit
– It is the segment of DNA between the sites of initiation and termination of transcription. It consists of 3 regions:
◦ A promoter: Binding site for RNA polymerase. Located towards 5′-end (upstream).
◦ Structural gene: The region between promoter and terminator where transcription takes place.
◦ A terminator: The site where transcription stops. Located towards 3′-end (downstream).
Transcription unit and gene
Gene is a functional unit of inheritance. It is the DNA sequence coding for an RNA (mRNA, rRNA or tRNA).
Cistron is a segment of DNA coding for a polypeptide during protein synthesis. It is the largest element of a gene.
Structural gene in a transcription unit is 2 types:
} Monocistronic structural genes (split genes): It is seen in eukaryotes. Here, coding sequences (exons or expressed sequences) are interrupted by introns (intervening sequences).
Exons appear in processed mRNA.
Introns do not appear in processed mRNA.
} Polycistronic structural genes: It is seen in prokaryotes. Here, there are no split genes.
Transcription in prokaryotes
In bacteria (Prokaryotes), synthesis of all types of RNA are catalysed by a single RNA polymerase. It has 3 steps:
} Initiation: Here, the enzyme RNA polymerase binds at the promoter site of DNA. This causes the local unwinding of the DNA double helix. An initiation factor (σ factor) present in RNA polymerase initiates the RNA synthesis.
} Elongation: RNA chain is synthesized in 5’-3’ direction. In this process, activated ribonucleoside triphosphates (ATP, GTP, UTP & CTP) are added. This is complementary to the base sequence in the DNA template.
} Termination: A termination factor (ρ factor) binds to the RNA polymerase and terminates the transcription.
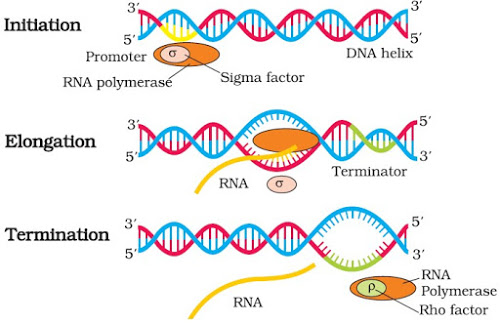 |
| Transcription in Bacteria |
In bacteria, transcription and translation can be coupled (translation begins before mRNA is fully transcribed) because
· mRNA requires no processing to become active.
· Transcription and translation take place in the same compartment (no separation of cytosol and nucleus).
Transcription in eukaryotes
In eukaryotes, there are 2 additional complexities:
1. There are 3 RNA polymerases:
· RNA polymerase I: Transcribes rRNAs (28S, 18S & 5.8S).
· RNA polymerase II: Transcribes the heterogeneous nuclear RNA (hnRNA). It is the precursor of mRNA.
· RNA polymerase III: Transcribes tRNA, 5S rRNA and snRNAs (small nuclear RNAs).
2. The primary transcripts (hnRNA) contain exons and introns and are non-functional. Hence introns must be removed. For this, it undergoes the following processes:
· Splicing: From hnRNA, introns are removed (by the spliceosome) and exons are spliced (joined) together.
· Capping: Here, a nucleotide methyl guanosine triphosphate (cap) is added to the 5’ end of hnRNA.
· Tailing (Polyadenylation): Here, adenylate residues (200-300) are added at 3’-end.
Now, it is the fully processed hnRNA, called mRNA.
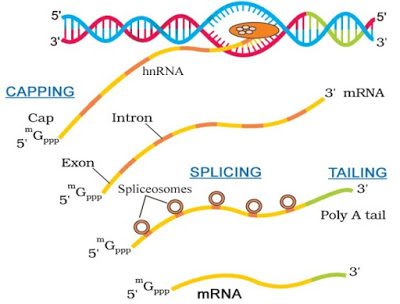 |
Transcription & Processing in Eukaryotes |
GENETIC CODE
§ It is the sequence of nucleotides (nitrogen bases) in mRNA that contains information for protein synthesis (translation).
§ The sequence of 3 bases determining a single amino acid is called codon.
§ George Gamow suggested that for coding 20 amino acids, the code should be made up of 3 nucleotides. Thus, there are 64 codons (43= 4 x 4 x 4).
§ Har Gobind Khorana developed the chemical method in synthesizing RNA molecules with defined combinations of bases (homopolymers & copolymers).
§ Marshall Nirenberg developed cell-free system for protein synthesis.
§ Severo Ochoa (polynucleotide phosphorylase) enzyme is
used to polymerize RNA with defined sequences in a template independent manner.
20 types of amino acids involved in translation
- Alanine (Ala)
- Arginine (Arg)
- Asparagine (Asn)
- Aspartic acid (Asp)
- Cystein (Cys)
- Glutamine (Gln)
- Glutamic acid (Glu)
- Glycine (Gly)
- Histidine (His)
- Isoleucine (Ile)
- Leucine (Leu)
- Lysine (Lys)
- Methionine (Met)
- Phenyl alanine (Phe)
- Proline (Pro)
- Serine (Ser)
- Threonine (Thr)
- Tryptophan (Trp)
- Tyrosine (Tyr)
- Valine (Val)
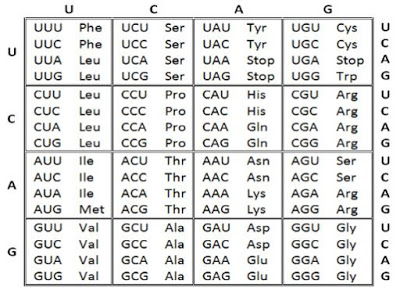
The codons for various amino acids
Salient features of genetic code
· Codon is triplet (three-letter code).
· 61 codons code for amino acids. 3 codons (UAA, UAG & UGA) do not code for any amino acids. They act as stop codons (Termination codons or non-sense codons).
· Genetic code is universal. E.g. From bacteria to human UUU codes for Phenylalanine. Some exceptions are found in mitochondrial codons, and in some protozoans.
· No punctuations b/w adjacent codons (comma less code). The codon is read in mRNA in a contiguous fashion.
· Genetic code is non-overlapping.
· An amino acid is coded by more than one codon (except AUG for methionine & UGG for tryptophan). Such codons are called degenerate codons.
· Genetic code is unambiguous and specific. i.e. one codon specifies only one amino acid.
· AUG has dual functions. It codes for Methionine and acts as initiator codon. In eukaryotes, methionine is the first amino acid and formyl methionine in prokaryotes.
Mutations and Genetic Code
– Relationship between genes & DNA are best understood by mutation studies. Deletions & rearrangements in a DNA may cause loss or gain of a gene and so a function.
– Insertion or deletion of one or two bases changes the reading frame from the point of insertion or deletion. It is called frame-shift insertion or deletion mutations.
– Insertion/ deletion of three or its multiple bases insert or delete one or multiple codon. The reading frame remains unaltered from that point onwards. Hence one or multiple amino acids are inserted /deleted.
– It proves that codon is a triplet and is read contiguously.
TYPES OF RNA
· mRNA (messenger RNA): Provide template for translation (protein synthesis).
· rRNA (ribosomal RNA): Structural & catalytic role during translation. E.g. 23S rRNA in bacteria acts as ribozyme.
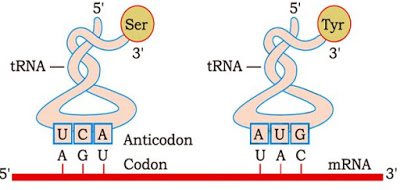
· tRNA (transfer RNA or sRNA or soluble RNA): Brings amino acids for protein synthesis and reads the genetic code.
Francis Crick postulated presence of an adapter molecule that can read the code and to link with amino acids.
tRNA is called adapter molecule because it has
· An Anticodon (NODOC) loop that has bases complementary to the codon.
· An amino acid acceptor end to which amino acid binds.
· Ribosome binding loop.
· Enzyme binding loop.
– For initiation, there is another tRNA called initiator tRNA.
– There are no tRNAs for stop codons.
– Secondary (2-D) structure of tRNA looks like a clover-leaf. 3-D structure looks like inverted ‘L’.
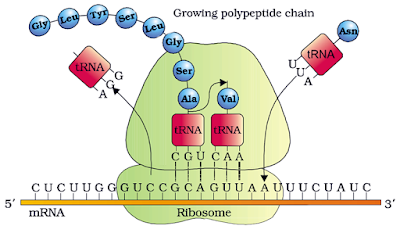
TRANSLATION (PROTEIN SYNTHESIS)
– It is the process of polymerisation of amino acids to form a polypeptide based on the sequence of codons in mRNA.
– It takes place in ribosomes. Ribosome consists of structural RNAs and about 80 types of proteins.
– Ribosome also acts as a catalyst (23S rRNA in bacteria is the enzyme- ribozyme) for the formation of peptide bond (peptidyl transferase enzyme in large subunit of ribosome).
– Translation includes 4 steps:
- Charging of tRNA
- Initiation
- Elongation
- Termination
1. Charging (aminoacylation) of tRNA
· Formation of peptide bond needs energy obtained from ATP.
· For this, amino acids are activated (amino acid + ATP) and linked to their cognate tRNA in presence of aminoacyl tRNA synthetase. Thus, the tRNA becomes charged.
2. Initiation
· In this, small subunit of ribosome binds to mRNA at the start codon (AUG).
· Now large subunit binds to small subunit to form initiation complex.
· Large subunit consists of aminoacyl tRNA binding site (A site) and peptidyl site (P site).
· The initiator tRNA (which carries methionine) binds on P site. Its anticodon (UAC) recognises start codon AUG.
3. Elongation
· Second aminoacyl tRNA binds to the A site of ribosome. Its anticodon binds to the second codon on the mRNA and a peptide bond is formed between first and second amino acids in presence of peptidyl transferase.
· First amino acid and its tRNA are broken. This tRNA is removed from P site and second tRNA from A site is pulled to P site along with mRNA. This is called translocation.
· These processes are repeated for other codons in mRNA.
· During translation, ribosome moves from codon to codon.
4. Termination
· When a release factor binds to stop codon, the translation terminates.
· The polypeptide and tRNA are released from the ribosomes.
· The ribosome dissociates into large and small subunits.
A group of ribosomes associated with a single mRNA for translation is called a polyribosome (polysomes).
An mRNA has additional sequences that are not translated (untranslated regions or UTR). UTRs are present at both 5’-end (before start codon) and 3’-end (after stop codon). They are required for efficient translation process.
REGULATION OF GENE EXPRESSION
In eukaryotes, gene expression occurs by following levels:
1. Transcriptional level (formation of primary transcript).
2. Processing level (splicing, capping etc.).
3. Transport of mRNA from nucleus to the cytoplasm.
4. Translational level (formation of a polypeptide).
The metabolic, physiological and environmental conditions regulate gene expression. E.g.
ú In E. coli, the beta-galactosidase enzyme hydrolyses lactose into galactose & glucose. In the absence of lactose, the synthesis of beta-galactosidase stops.
ú The development and differentiation of embryo into adult are a result of the expression of several set of genes.
If a substrate is added to growth medium of bacteria, a set of genes is switched on to metabolize it. It is called induction.
When a metabolite (product) is added, the genes to produce it are turned off. This is called repression.
OPERON CONCEPT
§ “Each metabolic reaction is controlled by a set of genes”
§ All the genes regulating a metabolic reaction constitute an Operon. E.g. lac operon, trp operon, ara operon, his operon, val operon etc.
Lac Operon in E. coli
– The operon controlling lactose metabolism.
– It is proposed by Francois Jacob & Jacque Monod.
It consists of
a) A regulatory or inhibitor (i) gene: Codes for repressor protein.
b) 3 structural genes:
i. z gene: Codes for b galactosidase. It hydrolyses lactose to galactose and glucose.
ii. y gene: Codes for permease. It increases permeability of the cell to b-galactosides (lactose).
iii. a gene: Codes for a transacetylase.
– Genes in the operon function together in the same or related metabolic pathway.
– If there is no lactose (inducer), lac operon remains switched off. The regulator gene synthesizes mRNA to produce repressor protein. This protein binds to the operator region and blocks RNA polymerase movement. So the structural genes are not expressed.
– If lactose or allolactose is provided in the growth medium, it is transported into E. coli cells by the action of permease. Lactose (inducer) binds with repressor protein. So repressor protein cannot bind to operator region. The operator region becomes free and induces the RNA polymerase to bind with promoter. Then transcription starts.
– Regulation of lac operon by repressor is called negative regulation.
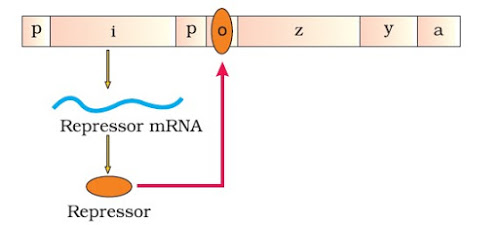 |
| In the absence of Inducer |
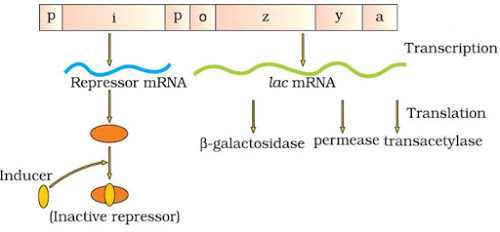 | |
|
HUMAN GENOME PROJECT (HGP)
· The entire DNA in the haploid set of chromosomes of an organism is called a Genome.
· In Human genome, DNA is packed in 23 chromosomes.
· Human genome contains about 3×109 bp.
· Human Genome Project (1990-2003) was the first mega project for the sequencing of nucleotides and mapping of all the genes in human genome.
· HGP was coordinated by U.S. Department of Energy and the National Institute of Health.
Goals of HGP
a. Identify all the estimated genes in human DNA.
b. Sequencing of 3 billion chemical base pairs of human DNA.
c. Store this information in databases.
d. Improve tools for data analysis.
e. Transfer related technologies to other sectors.
f. Address the ethical, legal and social issues (ELSI) that may arise from the project.
Methodologies of HGP: 2 major approaches.
- Expressed Sequence Tags (ESTs): Focused on identifying all the genes that are expressed as RNA.
- Sequence annotation: Sequencing whole set of genome containing all the coding & non-coding sequence and later assigning different regions in the sequence with functions.
Procedure of sequencing:
Isolate DNA from a cell → Convert into random fragments → Clone in a host (bacteria & yeast) using vectors (e.g. BAC & YAC) for amplification → Sequencing of fragments using Automated DNA sequencers (Frederick Sanger method) → Arrange the sequences based on overlapping regions→ Alignment of sequences using computer programs.
BAC= Bacterial Artificial Chromosomes
YAC= Yeast Artificial Chromosomes
- Sanger has also developed method for sequencing of amino acids in proteins.
- DNA is converted to fragments as there are technical limitations in sequencing very long pieces of DNA.
- HGP was closely associated with Bioinformatics.
- Bioinformatics: Application of computer science and information technology to the field of biology & medicine.
- Of the 24 chromosomes (22 autosomes and X & Y), the last sequenced one is chromosome 1 (May 2006).
- DNA sequencing also have been done in bacteria, yeast, Caenorhabditis elegans (a free living non-pathogenic nematode), Drosophila, plants (rice & Arabidopsis), etc.
Salient features of Human Genome
a. Human genome contains 3164.7 million nucleotide bases.
b. Total number of genes= about 30,000.
c. Average gene consists of 3000 bases, but sizes vary. Largest known human gene (dystrophin on X-chromosome) contains 2.4 million bases.
d. 99.9% nucleotide bases are same in all people. Only 0.1% (3×106 bp) difference makes every individual unique.
e. Functions of over 50% of discovered genes are unknown.
f. Chromosome I has most genes (2968) and Y has the fewest (231).
g. Less than 2% of the genome codes for proteins.
h. Very large portion of human genome is made of Repeated (repetitive) sequences. These are stretches of DNA sequences that are repeated many times. They have no direct coding functions. They shed light on chromosome structure, dynamics and evolution.
i. About 1.4 million locations have single-base DNA differences. They are called SNPs (Single nucleotide polymorphism or ‘snips’). This helps to find chromosomal locations for disease-associated sequences and tracing human history.
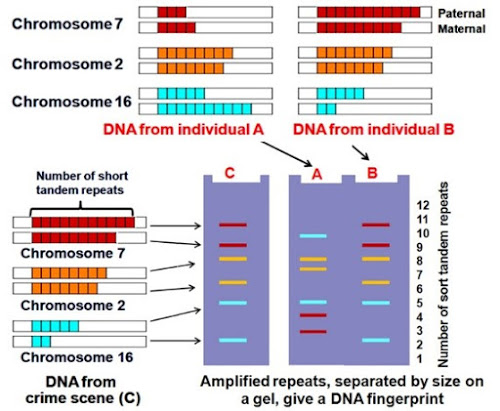
DNA FINGERPRINTING (DNA PROFILING)
- It is the technique to identify the similarities and differences of the DNA fragments of 2 individuals.
- It is developed by Alec Jeffreys (1985).
Basis of DNA fingerprinting
- DNA carries some non-coding repetitive sequences.
- Repetitive DNA can be separated from bulk genomic DNA as different peaks during density gradient centrifugation.
- The bulk DNA forms a major peak and the small peaks are called satellite DNA.
- Satellite DNA is classified as micro-satellites, mini-satellites etc. based on base composition (A:T rich or G:C rich), length of segment and number of repetitive units.
- A DNA sequence which is tandemly repeated in many copy numbers is called variable number tandem repeats (VNTR). It belongs to mini-satellite DNA.
- In a person, copy number varies in each chromosome.
- The two alleles (paternal and maternal) of a chromosome also contain different copy numbers of VNTR.
- VNTR is specific from person to person.
- The size of VNTR varies from 0.1 to 20 kb.
- Any difference in the nucleotide sequence (inheritable mutation) observed in a population is called DNA polymorphism (variation at genetic level).
- Polymorphism is higher in non-coding DNA sequence because mutations in these sequences may not affect an individual’s reproductive ability. These mutations accumulate generation to generation causing polymorphism.
- Polymorphisms have great role in evolution & speciation.
Steps of DNA fingerprinting (Southern Blotting Technique)
- Isolation of DNA (from any cells or blood stains, semen stains, saliva, hair roots, bone, skin etc.).
- Digestion of DNA by restriction endonucleases.
- Separation of DNA fragments by gel electrophoresis.
- Transferring (blotting) DNA fragments to synthetic membranes such as nitrocellulose or nylon.
- Hybridization using radioactive labelled VNTR probe.
- Detection of hybridized DNA by autoradiography.
The autoradiogram gives an image in the form of dark & light bands. It is called DNA fingerprint.
- DNA fingerprint differs in everyone except in monozygotic (identical) twins.
- The sensitivity of the technique can be increased by use of polymerase chain reaction (PCR). Therefore, DNA from a single cell is enough for DNA fingerprinting.
- Forensic tool to solve paternity, rape, murder etc.
- For the diagnosis of genetic diseases.
- To determine phylogenetic status of animals.
- To determine population and genetic diversities.
CBSE Class 12 Biology Important Questions Chapter 6 – Molecular Basis of Inheritance
1 Mark Questions
Chapter 6
Molecular Basis of Inheritance
1 Marks Questions
1. Name the factors for RNA polymerase enzyme which recognises the start and termination signals on DNA for transcription process in Bacteria.
Ans.Sigma (s) factor and Rho(p) factor)
2. Mention the function of non-histone protein.
Ans.Packaging of chromatin
3. During translation what role is performed by tRNA
Ans. (i) Structural role
(ii) Transfer of amino acid.
4. RNA viruses mutate and evolve faster than other viruses. Why?
Ans. -OH group is present on RNA, which is a reactive group so it is unstable and mutate faster.
5. Name the parts ‘X’ and ‘Y’ of the transcription unit given below.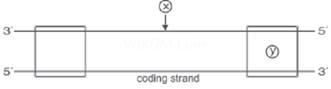
Ans. X – Template strand, Y – Terminator.
6. Mention the dual functions of AUG.
Ans. (i) Acts as initiation codon for protein synthesis
(ii) It codes for methionine.
7. Write the segment of RNA transcribed from the given DNA
3´ -A T G C A G T A C G T C G T A ‘5´- Template Strand
5´ – T A C G T C A T G C A G C A T ‘3´ – Coding Strand.
Ans. 5’- U A C G U C A U G C A G C A U – 3’ (In RNA ‘T’ is replaced by‘U’)
8.Name the process in which unwanted mRNA regions are removed & wanted regions are joined.
Ans.RNA splicing.
9.Give the initiation codon for protein synthesis. Name the amino acid it codes for?
Ans.Initiation codon – AUG & it code for methionine.
10.In which direction, the new strand of DNA synthesised during DNA replication.
Ans.5’→→ 3
11.What is the function of amino acyl tRNAsynthetase.
Ans.Amino acyl tRNAsynthetasecatalyses activation of amino and attachment of activated amino acids to the 3-end of specific tRNA molecule.
12.What is point mutation?
Ans.Mutation due to change in a single base pair in a DNA sequence is called point mutation.
13.Name the enzyme that joins the short pieces in the lagging strand during synthesis of DNA?
Ans.Ligase.
14.Name the enzyme which helps in formation of peptide bond?
Ans.Peptidyltransferase
15.Who experimentally prove that DNA replication is semi conservative.
Ans.Messelson&stahl.
16.What is a codon?
Ans.Triplet sequence of bases which codes for a single amino is called a codon.
17.Name the three non-sense codons?
Ans.UAA, UAG, UGA
18.What is the base pairing pattern of DNA?
Ans.In DNA, adenine always binds with thymine & cytosine always binds with Guanine.
19.Mention the dual functions of AUG?
Ans.AUG codes for amino acid methionine & also acts as an initiator codon.
2 Mark Questions
Chapter 6
Molecular Basis of Inheritance
2 Marks Questions
1. The process of termination during transcription in a prokaryotic cell is being represented here. Name the label a, b, c and d.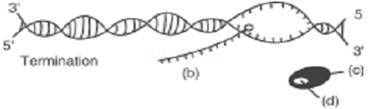
Ans. (a) DNA molecule
(b) mRNA transcript
(c) RNA polymers
(d) Rho factor
2. Complete the blanks a, b, c and d on the basis of Frederick Griffith Experiment.
S Strain →→ inject into mice →→ (a)
R strain →→ inject into mice →→ (b)
S strain (heat killed) →→ inject into mice →→ (c)
S strain (heat killed) + R strain (live) →→ inject into mice →→ (d)
Ans.(a) Mice die
(b) mice live
(c) mice live
(d) mice die
3. Give two reasons why both the strands of DNA are not copied during transcription.
Ans. (a) If both the strands of DNA are copied, two different RNAs(complementary to each other) and hence two different polypeptideswill produce; If a segment of DNA produces two polypeptides, thegenetic information machinery becomes complicated.
(b) The two complementary RNA molecules (produced simultaneously)would form a doublestranded RNA rather than getting translated intopolypeptides.
(c) RNA polymerase carries out polymerisation in 53direction andhence the DNA strand with 35 polarity acts as the template strand.(Any two)
4. Mention any two applications of DNA fingerprinting.
Ans.(i) To identify criminals in the forensic laboratory.
(ii) To determine the real or biological parents in case of disputes.
(iii) To identify racial groups to rewrite the biological evolution. (Any two)
5. State the 4 criteria which a molecule must fulfill to act as a genetic material.
Ans.(i) It should be able to generate its replica.
(ii) Should be chemically and structurally stable.
(iii) Should be able to express itself in the form of Mendelian characters.
(iv) Should provide the scope for slow changes (mutations) that are necessary for evolution.
6.“DNA polymerase plays a dual function during DNA replication” comment on statement?
Ans.DNA polymerase plays a dual function –it helps in synthesis of new strand & also helps in proof reading i.e replacement of RNA strands lay DNA fragments.
7.Three codons on mRNA are not recognised by tRNA what are they? What is the general term used for them what is their significance in protein synthesis?
Ans.UAG UAA & UGA are the three codons that are not recognised by tRNA these are known as stop codon or non-sense codon. Since these three codons are not recognised by any tRNA they help in termination of protein chain during translation.
8.Give two reasons why both the strands of DNA are not copied during DNA transcription?
Ans.I)If both the strands code for RNA two different RNA molecules & two different proteins wouldbe formed hence genetic machinery would become complicated
II) Since the two RNA molecules would be complementary to each other, they would wind togetherto form dsRNA without carrying out translation which means process of transcription would befutile
9.Why is it essential that tRNA binds to both amino acids & mRNA codon during protein synthesis?
Ans.It is essential that tRNA binds to both amino acids & mRNA codon because tRNA acts as an adapter molecule with picks up a specific activated aminoacid from the cytoplasm & transferred it to the ribosomal in the cytoplasm where proteins are synthesized. It attracts itself to ribosome with the sequence specified by mRNA & finally it transmits its amino acid to new polypeptide chain.
10.What is peptide bond? How is it formed?
Ans.Peptide bond is formed between carboxylic group (COOH) of first amino acid & amino group (-NH2) of second amino acid. This reaction is catalysed by peptidyltransferase
11.Explain what happens in frameshift mutation? Name one disease caused by the disorder?
Ans.Frameshift mutation is a type of mutation where addition or deletion of one or two bases changes the reading from the site of mutation, resulting in protein with different set of amino acid.
12.What do you mean by “Central Dogma of Molecular genetics?”
Ans.The central dogma of molecular genetics is the flow of genetic information from DNA to DNA through replication, DNA to mRNA through transcription & mRNA to proteins through translation.
ReplicationDNA →→ mRNA →→ proteins. transcription translation
13.Give two reasons why both the strands are not copied during transcription?
Ans.i) If both the strands codes for RNA, two different RNA molecules & two different proteins areformed hence genetic machinery would be complicated.
ii)Since two RNA molecules produced would be complementary to each other, they would wind together to form ds-RNA.
14.Why is human Genome project considered as mega project?
Ans.Human Genome project was called mega project for the following facts.
- The human genome has approximately 3.3 x 109bp, if the cost of sequencing is US 3perbp,theapproximatecostisaboutUS3perbp,theapproximatecostisaboutUS g billion.
- If the sequence obtained were to be stored in a typed form in books & if each page contained1000 letters & each book contained 1000 page than 3300 such books would be needed to store complete in formation
- The enormous quantity of data expected to be generated also necessitates the use of high speedcomputational devices for data storage, retrieval & analysis.
15.Why is DNA & not RNA is the genetic material in majority of organisms?
Ans.The -OH group in the nucleotides of RNA is much more reactive & makes RNA labile & easily degradable thus, DNA and not RNA acts as genetic material in majority of organisms.
16.Mention any four important characteristics of genetic code.
Ans.Genetic codon has following imp-features :-
- Each codon is a triplet consisting of three bases.
- Each codon codes for only one amino acid i.e. – unambiguous.
- Some amino acids are coded lay more than one codon ∴∴ said to be degenerative.
- Codons are read in a continuous manner in direction & have no punctuation.
17.Why it is that transcription & translation could be coupled in prokaryotic cell but not in eukaryotic cell?
Ans.In prokaryotes the mRNA synthesised does not require any processing to become active &both transcription & translation occurs in the same cytosol but In Eukaryotes, primary transcriptcontains both exon & intron & is subjected to a process called splicing where introns are removed &exons are joined in a definite order to form mRNA.
3 Mark Questions
Chapter 6
Molecular Basis of Inheritance
3 Marks Questions
1. Give six points of difference between DNA and RNA in their structure/chemistry and function.
Ans.
| DNA | RNA |
| (i) Double stranded molecules (ii) Thymine as pyrimidine base (iii) Pentose sugar is Deoxyribose (iv) Quite stable and not very reactive (v) Dictates the synthesis of Polypeptides (vi) Found in the nucleus. | (i) Single stranded molecules (ii) Uracil as pyrimidine base (iii) Sugar is Ribose (iv) 2´-OH makes it reactive (v) Perform their functions in protein synthesis. (vi) They are transported into the cytoplasm. |
2. Explain how does the hnRNA becomes the mRNA. OR Explain the process of splicing, capping and tailing which occur during transcription in Eukaryotes.
Ans.hnRNA is precursor of mRNA. It undergoes
(i) Splicing : Introns are removed and exons are joined together.
(ii) Capping : an unusual nucleotide (methyl guanosine triphosphate isadded to the 5´ end of hnRNA.
(iii) Adenylate residues (200-300) are added at 3´ end of hnRNA.
3. Name the three major types of RNAs, specifying the function of each inthe synthesis of polypeptide.
Ans. (i) mRNA-(Messenger RNA) : decides the sequence of amino acids.
(ii) tRNA-(Transfer RNA) : (a) Recognises the codon on mRNA (b) transport the aminoacid to the site of protein synthesis.
(iii) rRNA (Ribosomal RNA) : Plays the structural and catalytic role during translation.
4. Enlist the goals of Human genome project.
Ans. The Human Genome Project (HGP) is an international scientific research project with the goal of determining the sequence of chemical base pairs which make up human DNA, and of identifying and mapping all of the genes of the human genome from both a physical and functional standpoint
5. A tRNA is charged with the amino acid methionine.
(i) Give the anti-codon of this tRNA.
(ii) Write the Codon for methionine.
(iii) Name the enzyme responsible for binding of amino acid to tRNA.
Ans. (a) UAC (b) AUG (c) Amino-acyltRNAsynthetase.
6. Illustrate schematically the process of initiation, elongation and termination during transcription of a gene in a bacterium.
Ans.In bacteria, the mRNA provides the template, tRNA brings aminoacids and reads the genetic code, and rRNAs play structural and catalytic role during translation.
There is single DNA-dependent RNA polymerase that catalyses transcription of all types of RNA in bacteria.
RNA polymerase binds to promoter and initiates transcription (Initiation)
It somehow also facilitates opening of the helix and continues elongation
Once the polymerases reaches the terminator region, the nascent RNA falls off, so also the RNA polymerase. This results in termination
7.What is transformation? Describe Grifith’s experiment to show transformation? What did he prove from his experiment?
Ans.Transformation means change in genetic makeup of an individual. Fredrick Grifith conducted aseries of experiments on streptococcus pneumoniae. He observed two strains of this bacterium –one forming smooth colonies with capsule (s-type) & other forming rough colonies without capsule
(R-type)
(i) when live s-type cells are infected into mice, they produced pneumonia & mice dies.
(ii) When live R-type cells are infected into mice, disease was not produced did not appear.
(iii) When heat – killed S-type cells were infected into mice, the disease did not appear.
(iv) When heat killed S-type cells were mixed with live R-cells & infected into mice, the mice died.
He concluded that R-strain bacteria had somehow been transformed by heat –killed S-strain bacteria which must be due to transfer of genetic material
8.The base sequence on one strand of DNA is ATGTCTATA
(i) Give the base sequence of its complementary strand.
(ii) If an RNA strand is transcribed from this strand what would be the base sequence of RNA?
(iii) What holds these base pairs together?
Ans. (i) TACAGATAT.
(ii) UACAGAUAU
(iii) Hydrogen bonds hold these base pairs together. Adenine & thymine are bounded by twohydrogen bonds & cytosine & Guanine are bonded by three hydrogen bonds.
9.Two claimant fathers filed a case against a lady claiming to be the father of her only daughter. How could this case be settled identifying the real biological father?
Ans.This case to identify the real biological father could lee settled lay DNA – finger printingtechnique. In this technique :-
- first of all, DNA of the two claimants who has to be tested is isolated.
- Isolated DNA is then digested with suitable restriction enzyme & digest is subjected to gelelectrophoresis.
- The fragments of ds DNA are denatured to produce ss DNA by alkali treatment.
- The electrophoresed DNA is then transferred from get into a nitrocellulose filter paper where itis fixed.
- A known sequence of DNA is prepared called probe – DNA & is labelled with radioactive esotope32p & then probe is added to nitrocellulose paper.
- The nitrocellulose paper is photographed on X –ray film through auto radiography. The film isanalysed to determine the presence of hybrid nucleic acid.
Then, the DNA fingerprints of the two claimants is compared with the DNA fingerprint of the lady &her daughter, whosoever matches with each other would be declared as biological father of herdaughter.
10.The length of DNA in an eukaryotic cell is N 2.2 m How can such a huge DNA be packaged in a nucleus of micrometer in diameter.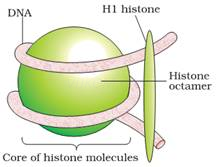
Ans.In eukaryotes, the DNA is wrapped around positivelycharged histone octamer into a structure called nucleosome. Atypical nucleosome consists of 200bp of DNA helix. Thenucleosomes are the repeating units that form chromatin fibres.
These chromatin fibres condense at metaphase stage of celldivision to form chromosomes. The packaging of chromatin at higher level requires additional set ofproteins called non-histone chromosomal proteins thus in nucleus, certain regions of the chromatinare loosely packed & they Stain lighter than the other region, these are called euchromatin. Theother region are lightly packed & they stain darker & are called heterochromatin
11.A tRNA is charged with amino acid methionine.
i) At what site in the ribosome will the tRNA bind?
ii) Give the anticodon of this tRNA?
iii) What is the mRNA codon for methionine?
iv) Name the enzyme responsible for this binding?
Ans. (i) P- site
(ii) UAC
(iii) AUG
(iv) Amino acyl tRNASynthetase
12.Describe the continuous & discontinuous Synthesis of DNA?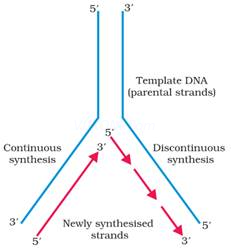
Ans.Synthesis of new strand of DNA takes place lay additionof fresh nucleotides to the 3 – OH group of the last nucleotideof the primer. This synthesis takes place in 5 direction&enzyme that catalyses this is DNA – polymerase
∴∴ synthesis of strand called leading strand iscontinuous.
The replication of second strand of the DNA molecule is
DISCONTINOUS on strand called lagging strand.
Primase initiates primer synthesis on strand near the fork. The RNA – primer thus formedprovides free for replication of single stranded region on lagging strand the newcomplementary strand is formed in small fragments of DNA called Okazaki fragments. It is calleddiscontinuous because it has to be initiated several times & every time an Okazaki fragment isproduced.
13.What are the three types of RNA & Mention their role in protein Synthesis?
Ans.There are three types of RNA :
- Messenger RNA (mRNA) :- It is a single – stranded RNA which brings the genetic information ofDNA transcribed on it for protein synthesis.
- Transfer RNA (tRNA) :- It has a clover leaf like structure which acts as an adapter moleculewhich contains an “anticodon loop” on one end that reads the code on one hand &” an amino acid acceptor end which binds to the specific amino acid on other hand.
- Ribosomal RNA (rRNA) :- Ribosomes provides the site for synthesis of protein &catalyse theformation of peptide bond.
14. Define bacterial transformation? Who proved it experimentally & how?
Ans. The transformation is a mode of exchange or transfer of genetic information betweenorganism or from one organism to another.
Fredrick Griffith tested the virulence of two strains of Diplococci to show transformation in thefollowing steps :-
- When S-III strains of bacteria are injected into mice. It developed pneumonia & died.
- When R-II strains are infected into mice, they did not develop pneumonia & survive.
- When heat – killed S-III strains of bacteria are injected into mice, No symptoms of pneumoniadevelops& mice remain healthy.
- When a mixture of heat – killed S-III strain & lives R-II strain is injected into mice, theydeveloped pneumonia & died.
From these results, Griffith concluded that the presence of heat – killed S-III bacteria must convertliving R-II type bacteria to type S-III so as to restore them the capacity for capsule formation. Thiswas called “BACTERIAL TRANSFORM ATION”
S strain →→ Inject into mice →→ Mice die
R strain →→Injct into mice →→ Mice live
S strain (heat-killed)→→Inject into mice →→ Mice live
S strain (heat-killed) + R strain (live) →→ Inject into mice →→Mice die
5 Marks Questions
Chapter 6
Molecular Basis of Inheritance
5 Marks Questions
1. What is meant by semi conservative replication? How did Meselson and Stahl prove it experimentally?
Ans.Meselson and Stahl, performed an experiment using E.coli to prove that DNA replication is semi conservative.
- They grew E.coli in a medium containing 15NH4Cl15NH4Cl.
- Then separated heavy DNA from normal (14N) by centrifugation in CsCl density gradient.
- The DNA extracted, after one generation of transfer from 15N medium to 14N medium, had an intermediate density.
-The DNA extracted after two generations consisted of equal amounts of light and hybrid DNA.
-They proved that DNA replicates in a semiconservative manner.
2. What does the lac operon consist of? How is the operator switch turned on and off in the expression of genes in this operon? Explain.
Ans.Lac Operon consists of the following :
- Structural genes : z, y, a which transcribe a polycistronic mRNA.
- gene ‘z’ codes for b-galactosidase
- gene‘y’ codes for permease.
- gene‘a’ codes for transacetylase.
- Promotor : The site where RNA polymerase binds for transcription.
- Operator : acts as a switch for the operon
- Repressor : It binds to the operator and prevents the RNAPolymerase from transcribing.
- Inducer : Lactose is the inducer that inactivates the repressor by binding to it.
- Allows an access for the RNA polymerase to the structural gene andtranscription.
3. What is an operon? Describe the major steps involved in an operon?
Ans.Operon is a group of controller & structural genes which controls the catabolism of the cell geneticallyeg lactose operon / lac operon.
(i) When inducer or lactose is absent :-
The lac regulator gene synthesize a repressor protein by transcription & translation. This repressor protein binds with operator site of lac operon & blocks RNA polymerase. Thus, RNA polymerase unable to transcribe mRNA & structural gene unable to translate enzyme B-galactosidase.
(ii) When inducer or lactose is present :_
The lac regulator gene transcribe mRNA &synthesise active lac repressor protein & at the same time lactose is converted into isomer allolactose. Allolactose binds to active lac repressor due to which it is converted to inactive repressor. This inactive repressor is released from operator site of lac operon & RNA polymerase binds to promoter & starts to transcribe mRNA & forms β-galactosidase are which converts lactose into glucose &galactose.
Thus, presence of lactose determines whether or not lac. Repressor is bound to operator & genes are expressed on not.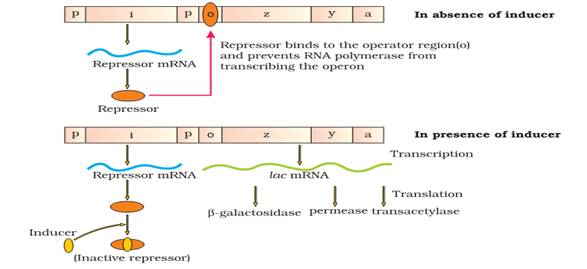
4.What do you mean semi conservative nature of DNA replication. Who proved it & how?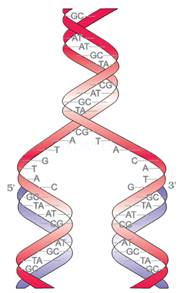
Ans.semiconservative nature of DNA replication suggested that during replication two strandswould separate & each acts as a template for the synthesis of new complementary strand so, thatafter complete replication, each DNA molecule would have one parental & one newly synthesizedstrand thus, half the information is conserved over generation. Mathew Messselson& Franklin stahl have performed an experiment using Escherichia coli to prove that DNA replication is Semiconservative. 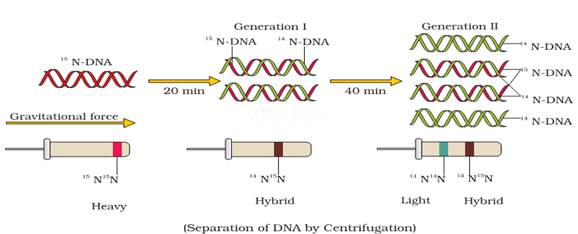
5.Where do transcription & translation takes place in a prokaryotic cell? Describe the three stepsinvolved in translation?
Ans.In a prokaryotic cell both transcription & translation occurs in cytoplasm. It consist offollowing steps :-
(i)ACTIVATION OF AMINO ACIDS :- amino acids are activated in the presence of ATP lay enzaminoacyltRNASynthetase.
(ii)BINDING OF ACTIVATED AMINOACID WITH tRNA :- Activated amino acids binds with specific tRNA to form charged tRNA .
(iii)INITIATION OF POLYPEPTIDE CHAIN :- Initiation codon is AUG which codes for methionine. Initiation codon of mRNA binds to p-site of ribosome with the help of initiation factors.
(iv)ELONGATION OF POLYPEPTIDE CHAIN :-
(a)Second activated amino acid along itstRNA reaches the ‘A’ site & binds to mRNA codon next to AUG.
(b)A peptide bond is formed betweentwo amino acid by peptidyl transferase.
(c) Ribosomes translocation mRNA in -direction due to which free tRNA slips away &peptidyltRNA reaches at P – site. Now third amino acid reaches at A – site & process continues.
(d)TERMINATION OF POLYPEPTIDE CHAIN :- When a termination codon (UAA, UAG, UGA) reaches at A- site translation terminates Since there is no specific tRNA for these codons.
(i)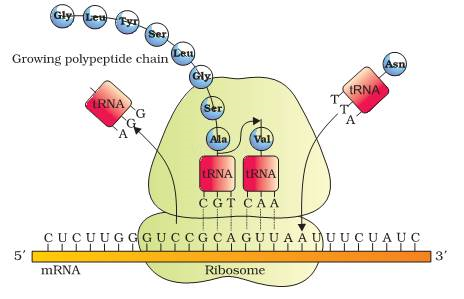
6.Who performed the blender experiment? What does this experiment prove? Describe the stepsfollowed in this experiment?
Ans.The proof for DNA as the genetic material came from the experiments of Harshey& chase whoworked with bacteriophage.
The bacteriophage on infection injects only the DNA into the bacterial cell & not the protein coat.
Bacterial cell treats the viral DNA as its own & subsequently manufactures more virus particles.
They grew some viruses on a medium that ‘contained radioactive Phosphorus & some other on medium that contained radioactive sulphur. Virus grown in the presence of radioactive phosphorus contained radioactive DNA but not proteins because DNA contains phosphorus. Similarly virus grown on radioactive sulfur contained radioactive protein because DNA does not contain sulfur.
Radioactive phages are allowed to infect E. coli bacteria & soon after infection the cultures weregently agitated in a blender to separate the adhering protein coat of virus from bacterial cell.It was found that when phage containing radioactive DNA was used to infect the bacteria itsradioactivity was found in bacterial cells indicating that DNA has been injected into bacterial cell so,the DNA is the genetic material & not proteins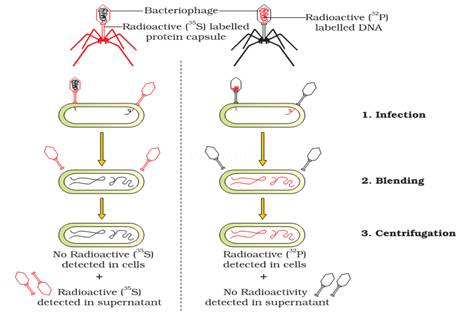
Molecular Basis of Inheritance Class 12 Biology MCQs
1. The DNA site where DNA-dependent RNA- polymerase binds for transcription, is called
(a) operator
(b) promotor
(c) regulator
(d) receptor
Answer
Answer: b
2. Operon model for regulation of transcription was proposed by
(а) Meselson and Stahl
(b) Jacob and Monod
(c) Watson and Crick
(d) Hershey and Chase
Answer
Answer: b
3. Eukaryotic RNA polymerase III catalyses the synthesis of
(a) mRNA
(b) rRNA
(c) hnRNA
(d) tRNA
Answer
Answer: d
4. The sequence of nitrogen bases in a # segment of a coding strand of DNA is ’ AATGCTTAGGCA. What will be the
sequence of nitrogen bases in the wRNA transcribed by it?
(a) UUA CGA AUC CGU
(b) AAU GCU AAC CGA
(c) AAU GCA AUC CGU
(d) AAU GCU UAG GCA
Answer
Answer: d
5. In the lac operon of E.coli, the i gene codes for
(a) inducer
(b) repressor
(c) lactase
(d) β-galactosidase
Answer
Answer: b
6. Which of the following sets of codons contains only termination codons?
(a) UAA, UGA, UAG
(b) UAA, UUU, UGG
(c) UAA, UAG, UAC
(d) UUU, UCC, UGG
Answer
Answer: a
7. The central dogma of molecular biology (genetic information flow) was modified by the discovery of
(a) RNA polymerase
(b) DNA ligase
(c) Reverse transcriptase
(d) DNA polymerase
Answer
Answer: c
8. The fact that a purine base always paired through hydrogen bonds with a pyrimidine base leads to, in the DNA double helix [NCERT Exemplar]
(a) the antiparallel nature
(b) the semiconservative nature
(c) uniform width throughout DNA
(d) uniform length in all DNA.
Answer
Answer: bc
9. The promoter site and the terminator site for transcription are located at [NCERT Exemplar]
(a) 3′ (downstream) end and 5′ (upstream) end, respectively of the transcription unit.
(b) 5′ (upstream) end and 3′ (downstream) end, respectively of the transcription unit.
(c) the 5′ (upstream) end.
(d) the 3′ (downstream) end.
Answer
Answer: b
10. The human chromosome with the highest and least number of genes in them are respectively [NCERT Exemplar]
(a) Chromosome 21 and Y
(b) Chromosome 1 and X
(c) Chromosome 1 and Y
(d) Chromosome X and Y
Answer
Answer: c
11. Discontinuous synthesis of DNA occurs in one strand, because [NCERT Exemplar]
(a) DNA molecule being synthesised is very long.
(b) DNA dependent DNA polymearse catalyses polymerisation only in one direction (5′ → 3′).
(c) it is a more efficient process.
(d) DNA ligase has to have a role.
Answer
Answer: b
12. Which of the following are the functions of RNA? [NCERT Exemplar]
(a) It is a carrier of genetic information from DNA to ribosomes synthesising polypeptides.
(b) It carries amino acids to ribosomes
(c) It is a constituent component of ribosomes
(d) All of the above.
Answer
Answer: d
13. In E.coli, the lac operon gets switched on when [NCERT Exemplar]
(a) lactose is present and it binds to the », repressor.
(b) repressor binds to operator.
(c) RNA polymerase binds to the operator.
(d) lactose is present and it binds to RNA polymerase.
Answer
Answer: a
14. The net electric charge on DNA and histone, is
(a) positive, negative
(b) negative, positive
(c) negative, negative
(d) positive, positive.
Answer
Answer: b
15. If the sequence of the nitrogen bases in the coding strand of DNA is 5’- ATGAATT-3’, the sequence of bases in the RNA transcribed by it will be ______ .
Answer/Explanation
Answer:
Explaination: 3’ AUGAAUU 5’
16. _____ step in transcription is catalysed by the enzyme DNA-dependent RNA polymerase.
Answer/Explanation
Answer:
Explaination: Elongation.
17. Lac operon shows the control of gene expression at the _____ level, in E.coli.
Answer/Explanation
Answer:
Explaination: Transcription.
18. The enzyme DNA polymerase catalyses the polymerisation of nucleotides in the ______ direction, for the lagging strand.
Answer/Explanation
Answer:
Explaination: 5′ → 3′
19. The last chromosome to be completely sequenced in the Human Genome Project (HGP) is ______ .
Answer/Explanation
Answer:
Explaination: Chromosome 1
20. RNA polymerase II in eukaryotes catalyses the transcription of ______ .
Answer/Explanation
Answer:
Explaination: hnRNA.
21. The presence of ______ group in every ribonucleotide makes RNA labile and reactive.
Answer/Explanation
Answer:
Explaination: 2′-OH.
22. Meselson and Stahl experimentally proved the _____ replication of DNA.
Answer/Explanation
Answer:
Explaination: Semiconservative.
23. During splicing in eukaryotes, the _____ are joined to from the RNA.
Answer/Explanation
Answer:
Explaination: Exons.
24. _____ factor functions as the initiation factor in the transcription of prokaryotes.
Answer/Explanation
Answer:
Explaination: Sigma (a)
25. Match the terms in Column I with those in Column II.
| Column I | Column II |
| A. Transcription | 1. A set of three bases on tRNA that is complementary to the bases of codon on mRNA. |
| B. Anticodon | 2. A unit of DNA that codes for a polypeptide. |
| C. Cistron | 3. Process of synthesis of polypeptide as dictated by mRNA. |
| D. Translation | 4. Process by which mRNA carries the information from nucleus to ribosomes. |
Answer/Explanation
Answer:
Explaination: A – 4, B – 1, C – 2, D – 3
26. Match the codons in Column I with the amino acids in Column II.
| Column I | Column II |
| A. UUU | 1. Termination |
| B. AUG | 2. Tyrosine |
| C. UAA | 3. Phenylalanine |
| D. AGU | 4. Methionine |
| E. UAC | 5. Serine |
Answer/Explanation
Answer:
Explaination: A – 3, B – 4, C – 1, D – 5, E – 2
27. Polycistronic mRNA is generally found in eukaryotes. [True/False]
Answer/Explanation
Answer:
Explaination: False.
28. The process of translation of mRNA begins, when the mRNA encounters the large subunit of ribosome [True/False]
Answer/Explanation
Answer:
Explaination: False.
29. VNTR belongs to a class of satellite DNA, called micro-satellite. [True/False]
Answer/Explanation
Answer:
Explaination: False.
30. If a double-stranded DNA contains 20% cytosine, it will have 20% guanine in it. [True/False]
Answer/Explanation
Answer:
Explaination: True.
31. Termination/Stop codons do not have any tRNAs [True/False]
Answer/Explanation
Answer:
Explaination: True.
Directions (Q32 to Q35): Mark the odd one in each of the following groups.
32. UAA, UGG, UAG, UGA
Answer/Explanation
Answer:
Explaination: UGG
33. 5S rRNA, snRNA, hnRNA, tRNA
Answer/Explanation
Answer:
Explaination: hnRNA
34. Har Gobind Khorana, Marshal Nirenberg, Severo Ochoa, Alec Jeffreys.
Answer/Explanation
Answer:
Explaination: Alec Jeffreys.
35. Promoter, Inducer, Operator, Terminator.
Answer/Explanation
Answer:
Explaination: Inducer.
36. Name the two types of nucleic acids in living systems.
Answer/Explanation
Answer:
Explaination: Deoxyribonucleic acid and ribonucleic acid.
37. How is the length of DNA usually defined?
Answer/Explanation
Answer:
Explaination:
Length of DNA is defined as the:
(i) number of nucleotides in single-stranded DNA, and
(ii) number of pairs of nucleotides (base pairs) in double-stranded DNA.
38. Name the specific components and the linkage between them that form deoxyadenosine. [Delhi 2013C]
Answer/Explanation
Answer:
Explaination: The nitrogenous base, adenine is linked to deoxyribose sugar through N-glycosidic linkage.
39. Name the specific components and the linkages between them that form deoxy- guanosine. [All India 2013C]
Answer/Explanation
Answer:
Explaination: The nitrogenous base guanine is linked to deoxyribose sugar by N-glycosidic linkage.
40. In which position is the phosphate group linked to a nucleoside? Name the linkage too.
Answer/Explanation
Answer:
Explaination:
– A phosphate group is linked to the 5′-OH of a nucleoside.
– Phosphoester linkage.
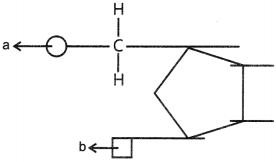
41. Name the components ‘a’ and ‘b’ in the nucleotide with a purine, given below:
Answer/Explanation
Answer:
Explaination:
a – Phosphate group,
b – Nitrogenous base
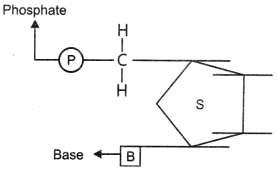
42. Mention the carbon positions to which the nitrogenous base and the phosphate molecule are respectively linked in the nucleotide given below:
Answer/Explanation
Answer:
Explaination:
– Nitrogenous base at the first carbon.
– Phosphate at the fifth carbon.
43. Mention the position of the ribonucleotide, where the OH group is present.
Answer/Explanation
Answer:
Explaination: The OH-group is present at the 2′-position.
44. Who discovered the nucleic acid DNA? What was it called then?
Answer/Explanation
Answer:
Explaination:
Frederich Meischer discovered DNA.
It was called nuclein.
45. The two strands of DNA have antiparallel polarity. What does it mean?
Answer/Explanation
Answer:
Explaination:
It means that one of the strands of DNA has
5′ → 3′ polarity and the other strand has
3′ → 5′ polarity.
46. If the base adenine constitutes 30 per cent of an isolated DNA fragment, then what is the expected percentage of the base cytosine in it? [Delhi 2011C; HOTS]
Answer/Explanation
Answer:
Explaination: 20 per cent.
47. How does the flow of genetic information in HIV7 deviate from the ‘central dogma’ proposed by Francis crick? [Foreign 2013]
Answer/Explanation
Answer:
Explaination: HIV shows reverse transcription, i.e. formation of DNA on RNA template.
48. How does HIV differ from a bacteriophage? [Delhi 201OC]
Answer/Explanation
Answer:
Explaination: HIV has RNA as its genome and shows reverse transcription, while bacteriophage has double-stranded DNA as its genome and no reverse transcription.
49. How is the length of DNA usually calculated?
Answer/Explanation
Answer:
Explaination: The length of DNA is calculated by multiplying the total number of base pairs with the distance between two consecutive base pairs, which is 0.34 nm or 0.34 × 10-9 m.
50. How may base pairs would a DNA segment of length 1.36 mm have? [Foreign 2017]
Answer/Explanation
Answer:
Explaination: It will have 1.36/0.34 × 10-9bp, i.e., 4.6 × 106bp.
51. What is a nucleoid?
Answer/Explanation
Answer:
Explaination: Nucleoid is the region in prokaryotic cells where DNA is organised as large loops held by some positively-charged proteins.
52. Name the positively charged protein around which the negatively charged DNA is wrapped. [AH India 2010C]
Answer/Explanation
Answer:
Explaination: Histone.
53. Name two basic amino acids that provide positive charge to histone proteins. [Delhi 2012C]
Answer/Explanation
Answer:
Explaination: Lysine and Arginine.
54. Write the role of histone proteins in packaging of DNA in eukaryotes. [Foreign 2017]
Answer/Explanation
Answer:
Explaination:
– Histone proteins are positively charged and are organized to from a unit of eight molecules, called a histone octamer,
– The negatively charged DNA molecule (of about 200 bp) wraps around the histone octamer to form a nucleosome, the repeating unit of chromatin.
55. Name the negatively charged and positively charged components of a nucleosome. [Delhi 2015C]
Answer/Explanation
Answer:
Explaination: Negatively charged component – DNA Positively charged component – histones.
56. Name the transcriptionally active region of chromatin in a nucleus. [Delhi 2015]
Answer/Explanation
Answer:
Explaination: Euchromatin.
57. Define transformation.
Answer/Explanation
Answer:
Explaination: Transformation is a phenomenon by which , the DNA isolated from one type of cell,
when introduced into another type, is able to bestow some of the properties of the former to the latter.
58. Write the conclusion Griffith arrived at, at the end of his experiments with Streptococcus pneumoniae. [All India 2017C]
Answer/Explanation
Answer:
Explaination:
– Griffith concluded that the R strain/non-virulent bacteria had been transformed y. into virulent form by the heat-killed S strain bacteria.
– It must be due to transfer of a transforming principle, i.e., genetic material from the heat-killed S strain bacteria.
59. What are bacteriophages?
Answer/Explanation
Answer:
Explaination: Bacteriophages are those viruses which infect the bacteria.
60. Why is RNA more reactive in comparison to DNA? [Delhi 2015CJ
Or
Why do RNA viruses undergo mutation and evolution faster than most of the DNA viruses? [HOTS]
Answer/Explanation
Answer:
Explaination: The 2′ OH-group in the nucleotides of RNA is a reactive group, that makes RNA labile 7- and easily degradable.
61. Write the scientific name of the plant on which Taylor et al performed their experiments.
Answer/Explanation
Answer:
Explaination: Vicia faba.
62. How long does the replication of human DNA take place?
Answer/Explanation
Answer:
Explaination: 38 minutes.
63. Name the enzyme and state its property that is responsible for continuous and discontinuous replication of the two strands of a DNA molecule. [Delhi 2013]
Answer/Explanation
Answer:
Explaination:
DNA-polymerase
– It polymerises the nucleotides only in 5′ → 3′ direction.
64. What is a replication fork?
Answer/Explanation
Answer:
Explaination: Replication fork is the Y-shaped structure formed when the double-stranded DNA is unwound up to a point during its replication.
65. Mention the direction in which:
(a) the leading strand is synthesised.
(b) discontinuous synthesis of DNA occurs.
Answer/Explanation
Answer:
Explaination:
(а) 5′ → 3′.
(b) 5′ → 3′.
66. Name the enzyme involved in the continuous replication of DNA strand. Mention the polarity of the template strand. [All India 2010]
Answer/Explanation
Answer:
Explaination:
DNA polymerase.
Template strand has 3′ → 5′ polarity.
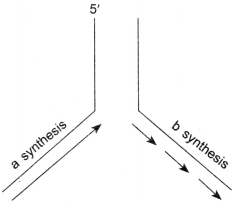
67. Name the types of synthesis ‘a’ and ‘b’ occurring in the replication fork of DNA as shown below:
Answer/Explanation
Answer:
Explaination:
‘a’ – Continuous synthesis
‘b’ – Discontinuous synthesis.
68. Name the enzyme that joins the small fragments of DNA of a lagging strand during DNA replication. [Delhi 2013C]
Answer/Explanation
Answer:
Explaination: DNA – ligase.
69. Why are vectors needed for replication of DNA during rDNA technology? [HOTS]
Answer/Explanation
Answer:
Explaination: The vector provides the origin of replication.
70. What will happen if DNA replication is not followed by cell division in a eukaryotic cell? [All India 2014C]
Answer/Explanation
Answer:
Explaination: It will lead to polyploidy, a condition where the cell comes to possess more than two sets of chromosomes.
71. Define transcription.
Answer/Explanation
Answer:
Explaination: Transcription is the process of copying the genetic information from one of the strands of DNA into RNA.
72. Name the enzyme and the direction in which it catalyses the polymerisation of ribonucleotides.
Answer/Explanation
Answer:
Explaination:
– DNA-dependent RNA-polymerase.
– 5′ → 3′ direction.
73. Mention one difference to distinguish an exon from an intron. [Foreign 2016]
Answer/Explanation
Answer:
Explaination:
– An exon is a coding sequence that forms part of RNA.
– An intron is a non-coding sequence that is removed during splicing and does not forms a part of RNA.
74. When and at what end does the ‘tailing’ of hnRNA take place?
Answer/Explanation
Answer:
Explaination: Tailing occurs after splicing at the 3′ end.
75. At which ends do ‘capping’ and ‘tailing’ of hnRNA occur, respectively?
Answer/Explanation
Answer:
Explaination: Capping at the 5′ end and tailing at the 3′ end.