Topic: 13.1
Chloroplasts carry out photosynthesis.
Fig. 1.1 shows some structural features of a chloroplast and some processes that occur within it.
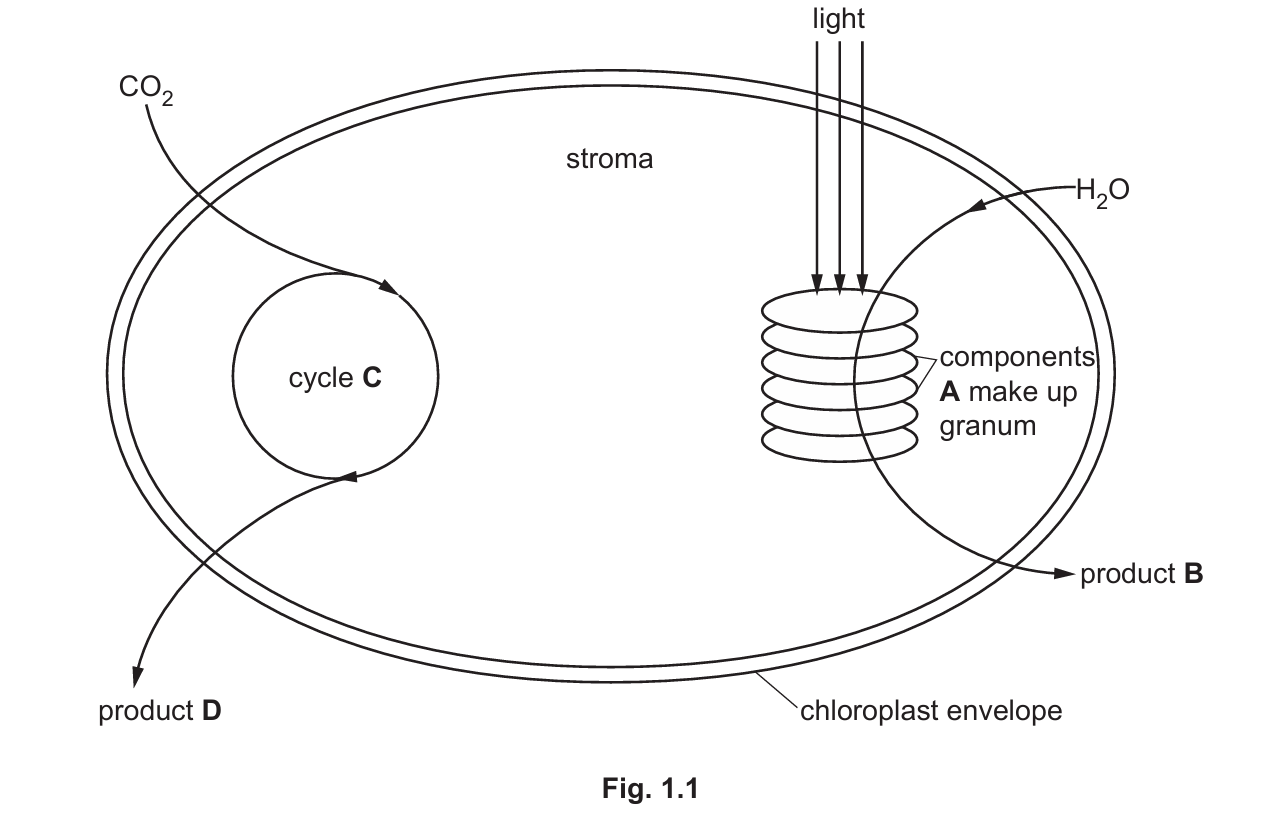
(a) (i) Identify the structures labelled A in Fig. 1.1.
(ii) Explain how the structure and appearance of the granum, and the components labelled A, relate to their function.
(b) (i) Identify the metabolic pathway labelled cycle C in Fig. 1.1.
(ii) Explain why pathway C is described as a cycle.
(c) (i) Identify the products of photosynthesis labelled B and D in Fig. 1.1.
(ii) Suggest and explain the importance of glucose and the product labelled B in Fig. 1.1 to ecosystems.
▶️ Answer/Explanation
(a)(i) thylakoid(s)
Explanation: The structures labelled A are the thylakoids, which are membrane-bound compartments inside chloroplasts where the light-dependent reactions of photosynthesis occur.
(a)(ii)
Explanation: The granum (stack of thylakoids) and thylakoids have several structural features that relate to their function:
- The stacked arrangement of thylakoids in grana creates a large surface area, which increases the efficiency of light absorption for photosynthesis.
- The thylakoid membranes contain photosynthetic pigments (like chlorophyll), photosystems, and electron carriers that are essential for the light-dependent reactions.
- The thylakoid lumen (inner space) accumulates protons (H⁺ ions) which creates a proton gradient used for ATP synthesis through chemiosmosis.
- The green appearance is due to chlorophyll pigments that absorb red and blue light for photosynthesis while reflecting green light.
(b)(i) Calvin (cycle)
Explanation: The cycle labelled C is the Calvin cycle, which is the light-independent stage of photosynthesis that fixes carbon dioxide into organic molecules.
(b)(ii)
Explanation: The Calvin cycle is described as a cycle because:
- It is a continuous process with no distinct start or end point – all intermediates are constantly present.
- The key substrate, ribulose bisphosphate (RuBP), is regenerated at the end of each turn of the cycle.
- It operates as a closed loop where the products of some reactions become the reactants for others.
(c)(i)
B: oxygen
D: sugar/glucose/triose phosphate/hexose/carbohydrate/starch
Explanation: During photosynthesis, water is split to release oxygen (B) as a byproduct, while carbon dioxide is fixed into organic compounds like glucose (D).
(c)(ii)
Explanation: The products of photosynthesis are crucial for ecosystems:
- Oxygen (B): Essential for aerobic respiration in most organisms, supporting oxidative phosphorylation which produces ATP.
- Glucose (D): Serves as the primary energy source for plants and organisms that consume plants, forming the basis of food chains.
- Together, these products facilitate energy flow through ecosystems, with glucose storing chemical energy that gets transferred between trophic levels.
- Oxygen also supports the respiratory needs of most organisms in the ecosystem.
Topic: 18.1
Biodiversity can be assessed at three different levels. One of these is the genetic variation within each species.
(a) Outline two other levels at which biodiversity can be measured.
To calculate the genetic variation that exists within a species, scientists:
- obtain DNA sequences from many individuals of one species
- count the number of nucleotides that differ when the sequences of two individuals are compared
- repeat this with different pairs of individuals.
This allows scientists to calculate the mean number of differences at every nucleotide position along the sequence (mean number of nucleotide differences per site).
(b) Explain why scientists use databases and computers to calculate the mean number of nucleotide differences per site.
(c) Table 2.1 shows the mean number of nucleotide differences per site of some species.
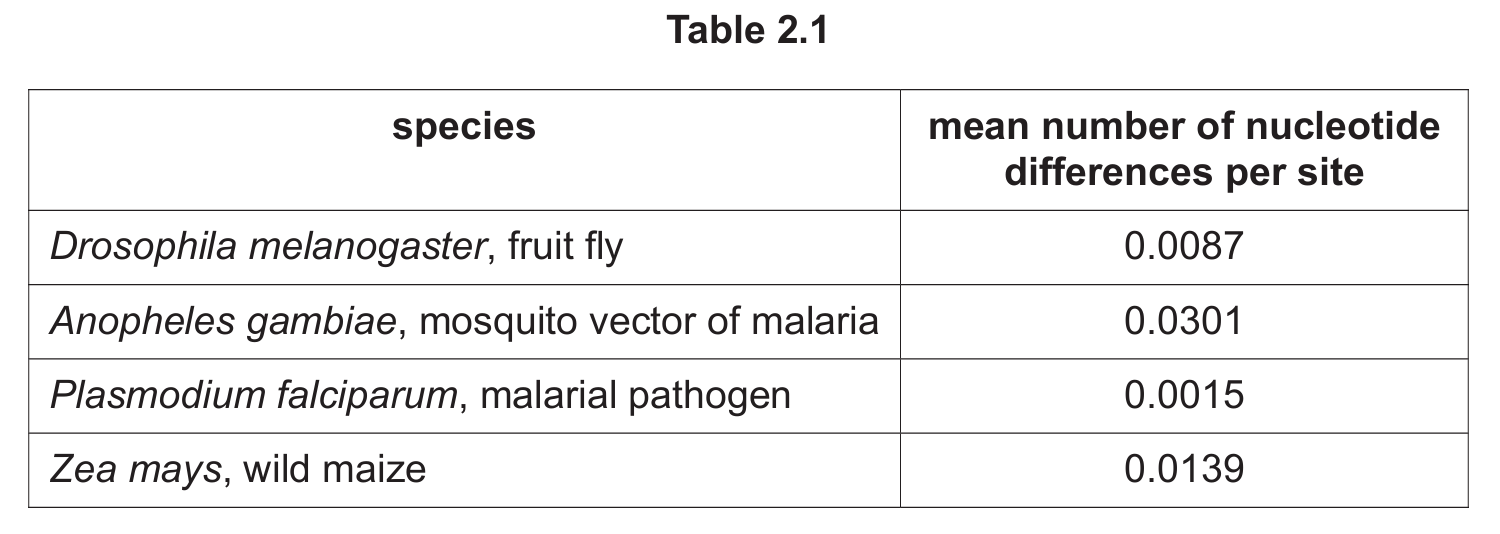
(i) State the genus name of the species that shows the most genetic variation.
(ii) State how many kingdoms of organisms are represented in Table 2.1.
(d) Genetic variation is considered important in the conservation of species. Low genetic variation is assumed to decrease the chance of the long-term survival of a species.
(i) Give reasons why low genetic variation may decrease the long-term survival of a species.
Fig. 2.1 shows how the International Union for the Conservation of Nature (IUCN) categorises species according to their conservation status.
Common species with the lowest conservation status (least risk of extinction) are categorised as Least Concern (LC).
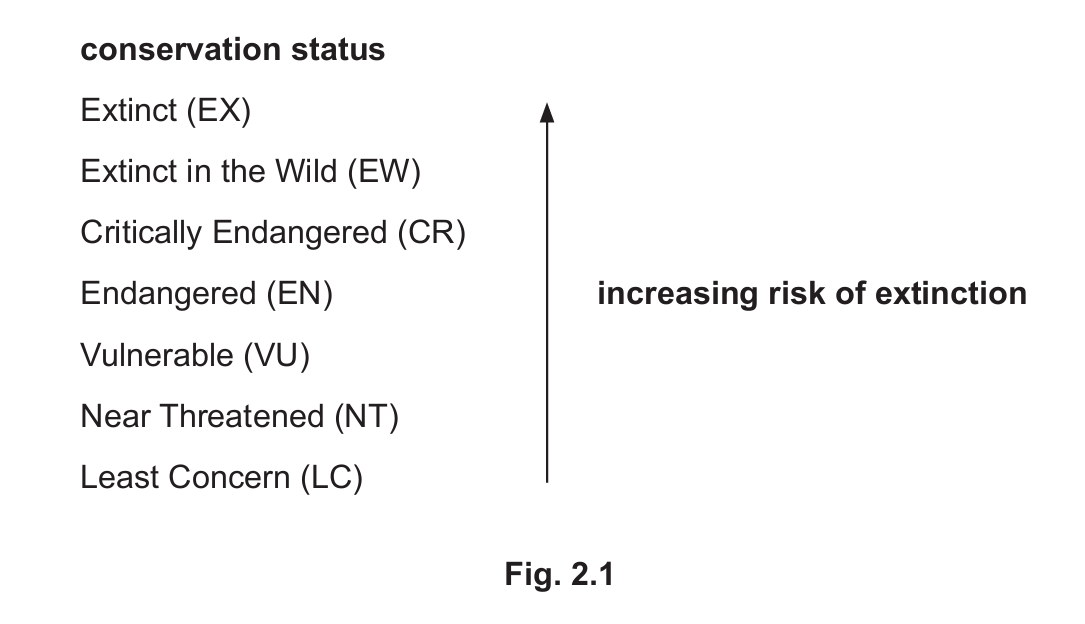
(ii) Question 2(d) states that ‘low genetic variation is assumed to decrease the chance of the long-term survival of a species’.
Predict the relationship between genetic variation and conservation status if this assumption is true.
Fig. 2.2 shows the mean number of nucleotide differences per site of some species and sub-species of mammal and their conservation status.
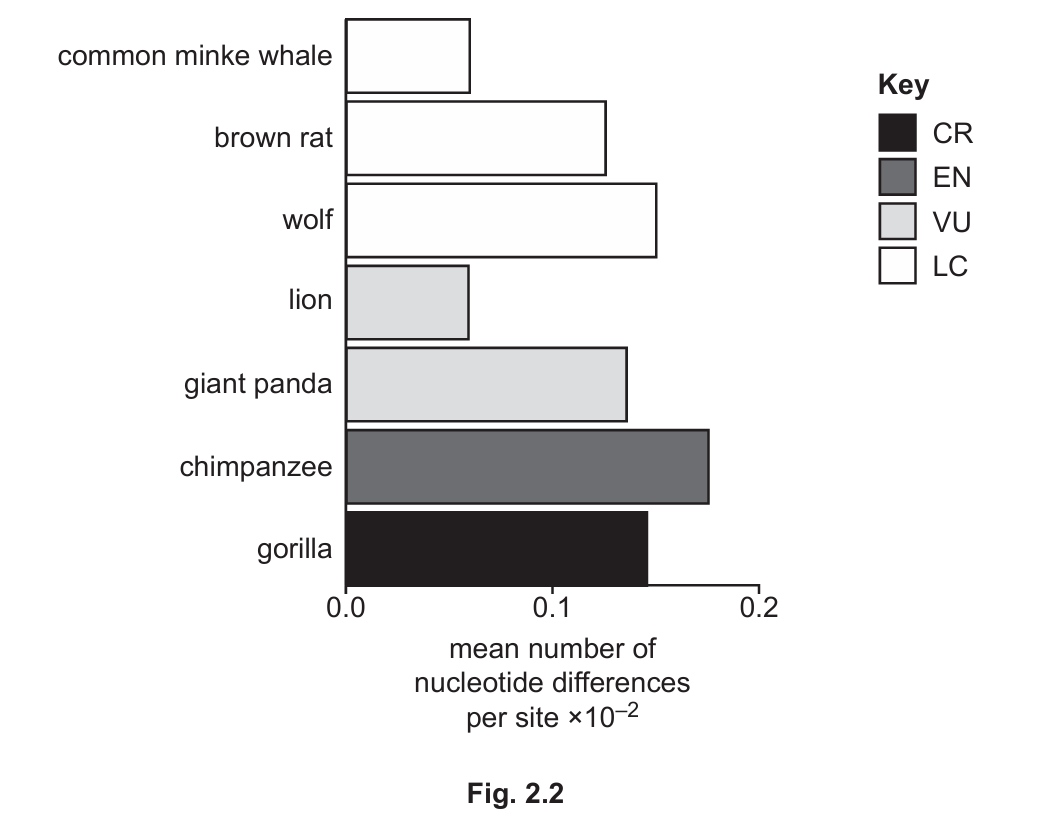
(iii) Assess whether the data in Fig. 2.2 provide support for the prediction you made in 2(d)(ii).
▶️ Answer/Explanation
(a)
1. Species diversity: The number of different species and their relative abundance in an ecosystem.
2. Ecosystem diversity: The variety of different habitats, communities, and ecological processes within a region.
Explanation: Biodiversity is typically measured at three levels: genetic diversity (within species), species diversity (between species), and ecosystem diversity (across habitats). The question asks for the two levels beyond genetic variation.
(b)
1. Databases can store vast amounts of genetic sequence data from multiple individuals.
2. Computers can quickly compare sequences and calculate differences between them.
3. Automated processing allows analysis of large datasets that would be impractical manually.
4. Digital tools enable statistical analysis and visualization of genetic variation patterns.
Explanation: Modern genetic studies generate enormous amounts of data. Computers and databases are essential for efficiently storing, organizing, and analyzing this information. They allow scientists to handle complex calculations and comparisons across entire genomes that would be impossible to do manually.
(c)(i) Anopheles
Explanation: The mosquito Anopheles gambiae shows the highest mean nucleotide difference (0.0301), indicating the most genetic variation among the listed species.
(c)(ii) Three kingdoms
Explanation: The table represents:
- Animalia (Drosophila, Anopheles)
- Protista (Plasmodium)
- Plantae (Zea mays)
(d)(i)
1. Reduced ability to adapt to environmental changes due to limited genetic options.
2. Increased susceptibility to diseases as pathogens can more easily exploit genetic uniformity.
3. Higher risk of inbreeding depression leading to harmful recessive traits being expressed.
Explanation: Genetic variation provides the raw material for natural selection. Populations with low genetic diversity have fewer options when facing environmental challenges, making them more vulnerable to extinction.
(d)(ii) Species with lower genetic variation would have higher conservation status (more endangered).
Explanation: If low genetic variation decreases survival chances, we’d expect such species to be more threatened (higher on IUCN scale) than genetically diverse species.
(d)(iii)
The data partially supports but also contradicts the prediction:
1. Supports: The giant panda (CR) has relatively low genetic variation (≈0.05 × 10⁻²).
2. Contradicts: The chimpanzee (EN) shows the highest variation (≈0.18 × 10⁻²).
3. The common minke whale (LC) has the lowest variation but least concern status.
Explanation: While some endangered species show low genetic diversity (supporting the hypothesis), there are exceptions where highly endangered species maintain considerable variation. This suggests genetic diversity is just one factor affecting conservation status.
Topic: 16.2
There are more than 600 plant species in the genus Ipomoea. Many species are grown for their attractive flowers, and some species are used as crop plants.
(a) Fig. 3.1 shows Ipomoea purpurea, the common morning glory.

The gene that determines flower colour in I. purpurea has two alleles:
- a dominant allele that results in purple flowers
- a recessive allele that results in red flowers.
A student recorded the flower colour of all the I. purpurea plants in a field. The student recorded 660 plants with purple flowers and 440 plants with red flowers.
Assuming the Hardy-Weinberg principle applies to this population, calculate the number of plants in the field that are heterozygous.
Use the equations:
\[ p + q = 1 \]
\[ p^2 + 2pq + q^2 = 1 \]
Show your working and give your answer to the nearest whole number.
(b) The Japanese morning glory, I. nil, has over 20 different flower colour phenotypes, including shades of blue, purple, red and pink.
The flower colour of I. nil is controlled by at least four genes. The flower colour can change gradually after the flowers open each morning and can change with fluctuations in the carbon dioxide concentration of the surrounding air.
A student concluded that the flower colour phenotype in I. nil shows continuous variation.
Suggest two reasons why the student made this conclusion.
(c) Scientists investigated the response of stomata to changing carbon dioxide (CO2) concentrations in the beach morning glory, I. pes-caprae.
The scientists placed I. pes-caprae plants in chambers. They measured the width of open stomata (stomatal apertures) after the plants had been exposed to different CO2 concentrations for 40 minutes. Light intensity and temperature were kept constant.
The relationship between CO2 concentration and the mean width of stomatal apertures is shown in Fig. 3.2.
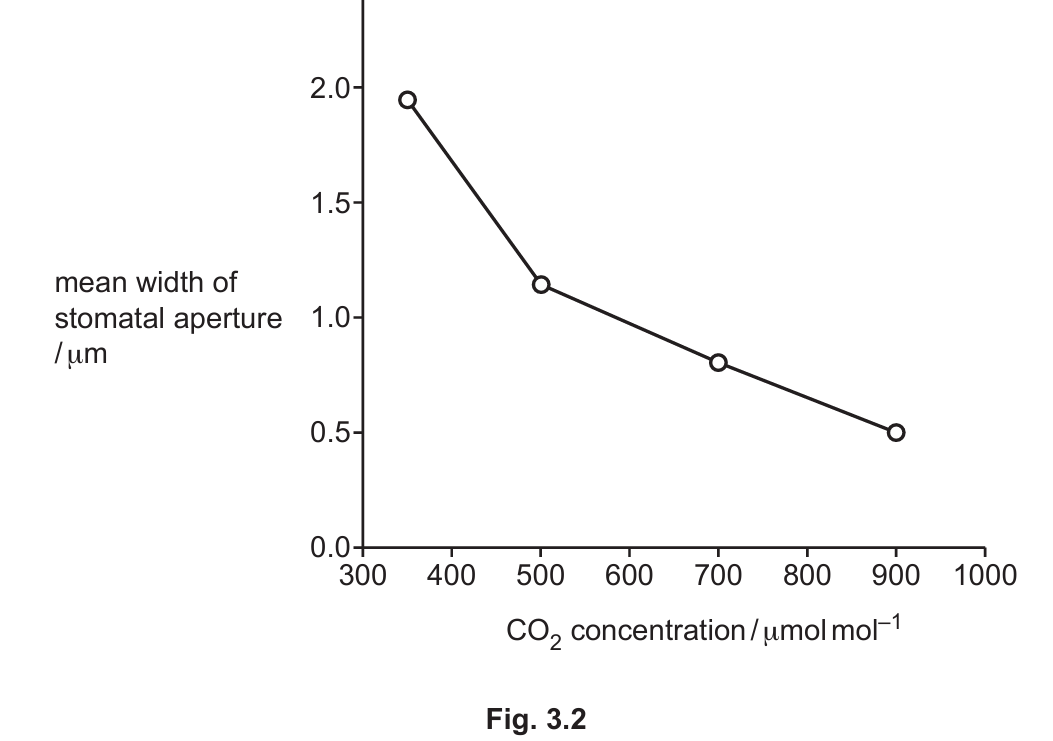
(i) In 2016, a study measured the atmospheric CO2 concentration as 400 μmol mol-1. In the future, climate change may reduce water availability and increase atmospheric CO2 concentrations in some habitats.
Suggest how the stomatal response shown in Fig. 3.2 would allow I. pes-caprae to survive the effects of climate change.
(ii) Under certain conditions, the closure of stomata is controlled by abscisic acid.
Describe how abscisic acid causes the closure of stomata.
(d) Scientists are researching whether abscisic acid can be used in crop treatment to increase yield. Evidence suggests that abscisic acid modifies the effect of auxin on elongation growth in plants.
(i) Scientists investigated the effect of different concentrations of abscisic acid on root elongation in seedlings of thale cress, Arabidopsis thaliana.
The seedlings were divided into four groups:
- a control group (0.0 μmol abscisic acid)
- three experimental groups, each treated with a different concentration of abscisic acid: 0.1 μmol, 1.0 μmol, or 10.0 μmol.
For each group of seedlings, root length was measured for six days during treatment. The rate of root elongation was calculated each day.
The results are shown in Fig. 3.3.
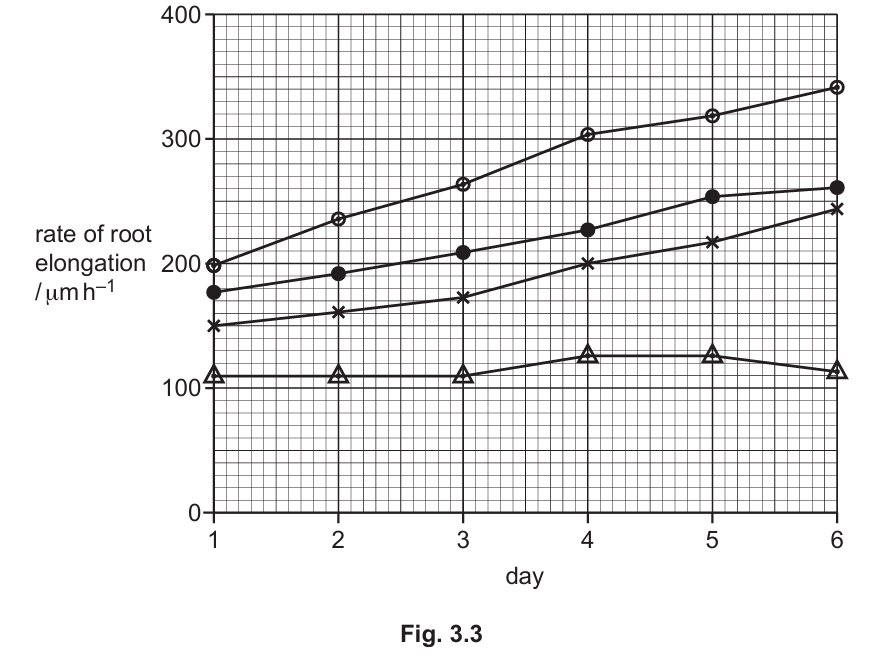

With reference to Fig. 3.3, describe the effect of treatment with abscisic acid on the rate of root elongation.
(ii) The passage outlines the role of auxin in elongation growth in plants.
Complete the passage by using the most appropriate scientific terms.
The binding of auxin to receptors causes …… to be pumped into cell walls.
This activates proteins called expansins, which disrupt the links between …… microfibrils. The cell walls are then able to expand.
▶️ Answer/Explanation
(a)
1. Total plants = 660 (purple) + 440 (red) = 1100
2. Red flowers are recessive (q2): q2 = 440/1100 = 0.4
3. q = √0.4 = 0.632
4. p = 1 – q = 1 – 0.632 = 0.368
5. Heterozygous frequency (2pq) = 2 × 0.632 × 0.368 = 0.465
6. Number of heterozygous plants = 0.465 × 1100 = 511.5 ≈ 512
Final answer: 511/512
Explanation: Using the Hardy-Weinberg principle, we first calculate the allele frequencies. The recessive phenotype (red flowers) gives us q2, from which we find q. Then p is calculated as 1-q. The heterozygous frequency is 2pq, which when multiplied by the total population gives the number of heterozygous plants.
(b)
1. The flower colors form a continuous range with many intermediate shades rather than distinct categories.
2. The phenotype is controlled by multiple genes (at least four), which typically results in continuous variation.
3. The phenotype is influenced by environmental factors (CO2 concentration and time of day).
Explanation: Continuous variation typically shows these characteristics: a range of phenotypes without clear categories, polygenic inheritance, and environmental influence on the phenotype.
(c)(i)
1. As CO2 concentration increases, stomatal apertures decrease (as shown in Fig. 3.2).
2. This reduces water loss through transpiration when water availability is low.
3. The plant can therefore conserve water while still obtaining sufficient CO2 for photosynthesis.
Explanation: The graph shows an inverse relationship between CO2 concentration and stomatal aperture. This adaptive response helps plants survive in drier conditions by reducing water loss while maintaining photosynthesis.
(c)(ii)
1. Abscisic acid (ABA) binds to receptors on guard cell membranes.
2. This triggers calcium ions (Ca2+) to enter the cytoplasm, acting as secondary messengers.
3. Potassium ions (K+) leave the guard cells.
4. Water follows by osmosis, moving out of the guard cells.
5. The guard cells become flaccid, closing the stomatal pore.
Explanation: ABA is a stress hormone that helps plants conserve water during drought conditions. The ion movements change the water potential in guard cells, leading to water loss and stomatal closure.
(d)(i)
1. At 0.1 μmol concentration, abscisic acid slightly increases the rate of root elongation compared to the control.
2. At higher concentrations (1.0 μmol and 10.0 μmol), abscisic acid decreases the rate of root elongation.
3. The 10.0 μmol treatment shows the most significant inhibition, with the rate remaining nearly constant over the six days.
Explanation: The graph shows a hormetic response – low concentrations stimulate growth while higher concentrations inhibit it. This suggests ABA has concentration-dependent effects on root growth.
(d)(ii)
1. The binding of auxin to receptors causes protons/H+ ions to be pumped into cell walls.
2. This activates proteins called expansins, which disrupt the links between cellulose microfibrils. The cell walls are then able to expand.
Explanation: Auxin promotes cell elongation through the acid growth hypothesis. Proton pumps acidify the cell wall, activating expansins that loosen the cellulose network, allowing turgor pressure to expand the cell.
Topic: 19.2
The potato plant, Solanum tuberosum, is an important food crop. Crop yield is reduced if the leaves of the plant are eaten by the larvae (immature stages) of the Colorado beetle, Leptinotarsa decemlineata.
Crop scientists used recombinant DNA technology to create two genetically modified (GM) varieties of potato plant. These plants produce proteins that are poisonous to insects.
- GM potato variety A contains two new genes, SN and Bt.
- GM potato variety B contains two new genes, SN and OClI.
The new varieties were tested by having a constant number of Colorado beetle larvae introduced to the plants at time 0 hours. The number of larvae that were alive after 24, 48 and 72 hours was recorded. The percentage of the larvae that had died in each time interval was calculated. This was repeated for potato plants that had not been genetically modified (non-GM).
Table 4.1 shows the percentage of Colorado beetle larvae that had died on the GM potato plant varieties and on non-GM potato plants.
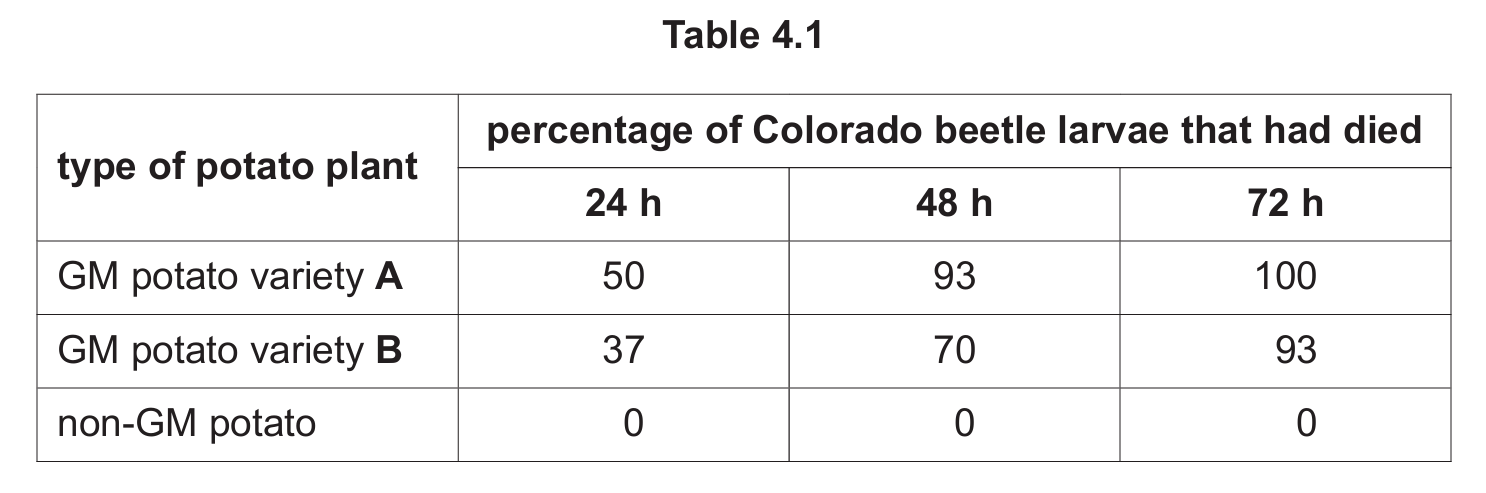
(a) (i) Suggest what is meant by recombinant DNA technology.
(ii) Suggest why the scientists created two different types of GM potato plant.
(iii) State why the scientists also performed the test on non-GM potato plants.
(b) Discuss how the results in Table 4.1 provide information that could help to solve the global demand for food.
▶️ Answer/Explanation
(a) (i) Recombinant DNA technology involves combining DNA from two different sources/species using enzymes or vectors to create a genetically modified organism (GMO) or transgenic organism.
Explanation: This technology allows scientists to take beneficial genes from one organism and insert them into another. For example, the Bt gene comes from the bacterium Bacillus thuringiensis and makes proteins toxic to certain insects. The process typically uses restriction enzymes to cut DNA and ligase enzymes to join DNA fragments, often with vectors like plasmids to transfer genes between organisms.
(a) (ii) To test/compare the effectiveness of the different genes/proteins (Bt and OClI) or plants/varieties (A and B) to see which works better.
Explanation: Creating two different varieties allows scientists to compare their effectiveness against the Colorado beetle. Variety A with SN and Bt genes might work differently than variety B with SN and OClI genes. This comparison helps determine which genetic combination provides better pest resistance, which is important for developing effective GM crops.
(a) (iii) To compare/provide a baseline or as a control.
Explanation: Non-GM plants serve as a control group in the experiment. They show what happens naturally without any genetic modification. This baseline is crucial for demonstrating that any observed effects (larval death) are indeed due to the genetic modifications and not other factors. It validates the experiment’s results by showing that without the GM traits, no larvae die.
(b) Both GM varieties A and B kill larvae, with variety A being more effective (killing 100% by 72 hours compared to 93% for variety B). GM technology increases potato yield by protecting plants from pests. Variety A, being most effective (killing larvae fastest and most completely), would provide the most food and should be prioritized for cultivation to help meet global food demand.
Explanation: The results clearly show that both GM varieties are effective against the Colorado beetle larvae, while non-GM plants offer no protection. Variety A stands out as particularly effective, killing all larvae within 72 hours. This pest protection directly translates to higher crop yields because undamaged plants can grow better and produce more potatoes. With global food demand increasing, such GM technologies that reduce crop losses to pests can significantly boost food production. The data suggests that adopting variety A could maximize this benefit, as it provides the most complete and fastest-acting pest control among the tested varieties. This approach aligns with sustainable agriculture goals by potentially reducing the need for chemical pesticides while increasing food production.
Topic: 12.2
(a) The respiratory quotient (RQ) values for different respiratory substrates can be calculated.
Table 5.1 shows:
- the formulae of four respiratory substrates
- the number of oxygen molecules (O2) needed to completely respire each substrate to carbon dioxide and water.
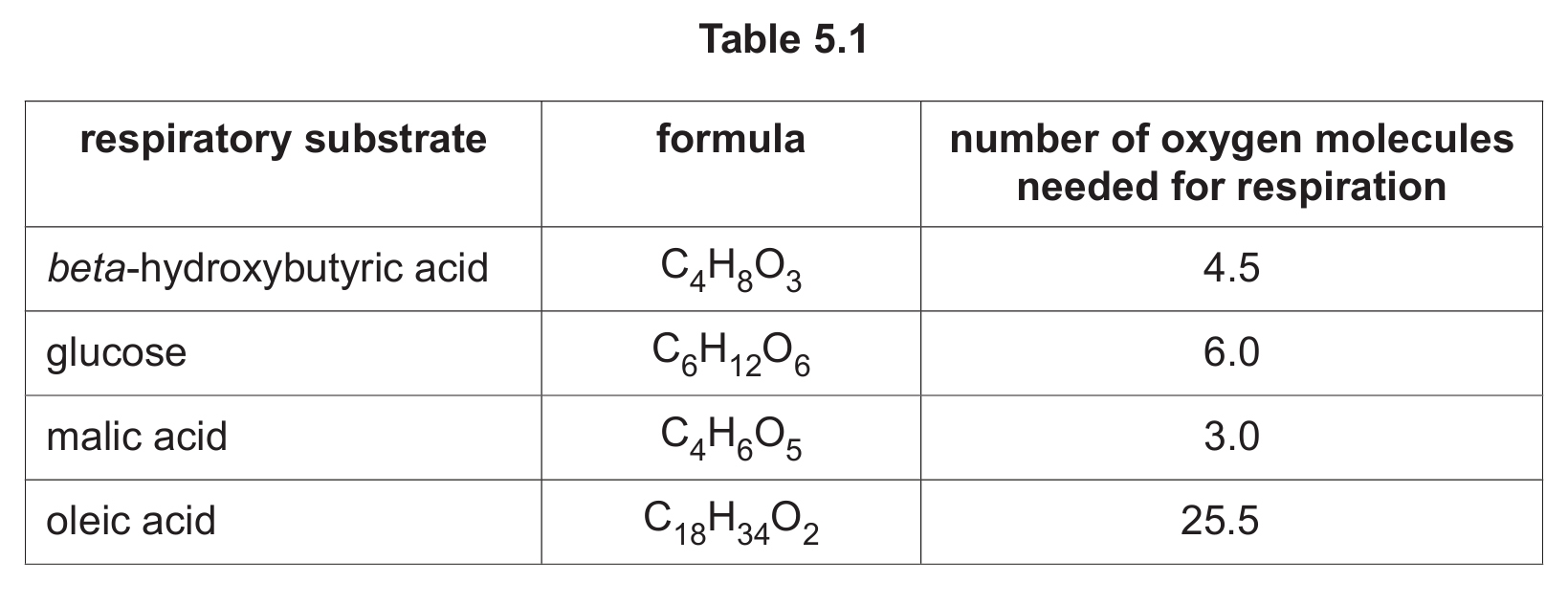
Using Table 5.1, name the respiratory substrate with the highest RQ and the respiratory substrate with the lowest RQ.
(b) In anaerobic conditions, the production of ATP in mammals and yeast involves glycolysis and fermentation.
Describe the similarities and differences between fermentation in mammals and in yeast.
▶️ Answer/Explanation
(a)
respiratory substrate with the highest RQ: malic acid
respiratory substrate with the lowest RQ: oleic acid
Explanation:
The respiratory quotient (RQ) is calculated as the ratio of CO2 produced to O2 consumed during respiration. For complete oxidation:
1. Malic acid (C4H6O5) requires 3 O2 molecules and produces 4 CO2 molecules, giving RQ = 4/3 ≈ 1.33 (highest).
2. Oleic acid (C18H34O2) is a fatty acid requiring 25.5 O2 molecules and producing 18 CO2 molecules, giving RQ = 18/25.5 ≈ 0.7 (lowest).
3. Glucose (C6H12O6) has RQ = 1 (balanced equation).
4. Beta-hydroxybutyric acid (C4H8O3) is a ketone body with RQ ≈ 0.89.
(b)
Similarities:
- Both processes use pyruvate from glycolysis as the starting molecule
- Both occur in the cytoplasm/cytosol of cells
- Both regenerate NAD+ from NADH to allow glycolysis to continue
- Both involve redox reactions (oxidation of NADH and reduction of organic molecules)
Differences:
- End products: Mammals produce lactate (lactic acid) while yeast produces ethanol and CO2
- Steps: Mammalian fermentation is a single-step reduction of pyruvate to lactate, while yeast fermentation has two steps (pyruvate → acetaldehyde → ethanol)
- Gas production: Yeast releases CO2 as a byproduct, while mammalian fermentation doesn’t produce gas
- Reversibility: Lactate fermentation is reversible (lactate can be converted back to pyruvate), while ethanol fermentation is irreversible
- Enzymes: Different enzymes are involved – lactate dehydrogenase in mammals vs pyruvate decarboxylase and alcohol dehydrogenase in yeast
Explanation:
Fermentation pathways diverge after glycolysis based on the organism’s enzymatic capabilities. Mammals convert pyruvate to lactate to regenerate NAD+ quickly during oxygen debt (e.g., intense exercise). Yeast, being fungi, have evolved the ability to decarboxylate pyruvate to acetaldehyde and then reduce it to ethanol, which is less toxic to their cells than lactate would be. The CO2 production in yeast fermentation is economically important in baking and brewing industries.
Topic: 17.1
The tiger barb, Puntigrus tetrazona, is a South American fish that is popular worldwide as an aquarium fish. Fig. 6.1 shows the appearance (phenotype) of a normal (wild-type) tiger barb.

– Tiger barbs that show a wild-type phenotype are gold with black stripes.
– Tiger barbs that show an albino phenotype are gold with white stripes.
– In 2012, a fish breeder discovered a tiger barb with a new, transparent, phenotype. This fish had a transparent body and black stripes.
The fish breeder crossed the tiger barb showing the new transparent phenotype with a tiger barb showing the albino phenotype.
All the F1 offspring were wild-type. These F1 offspring were crossed with each other.
Table 6.1 shows the phenotypes obtained in the F2 generation and the number of fish showing each phenotype.
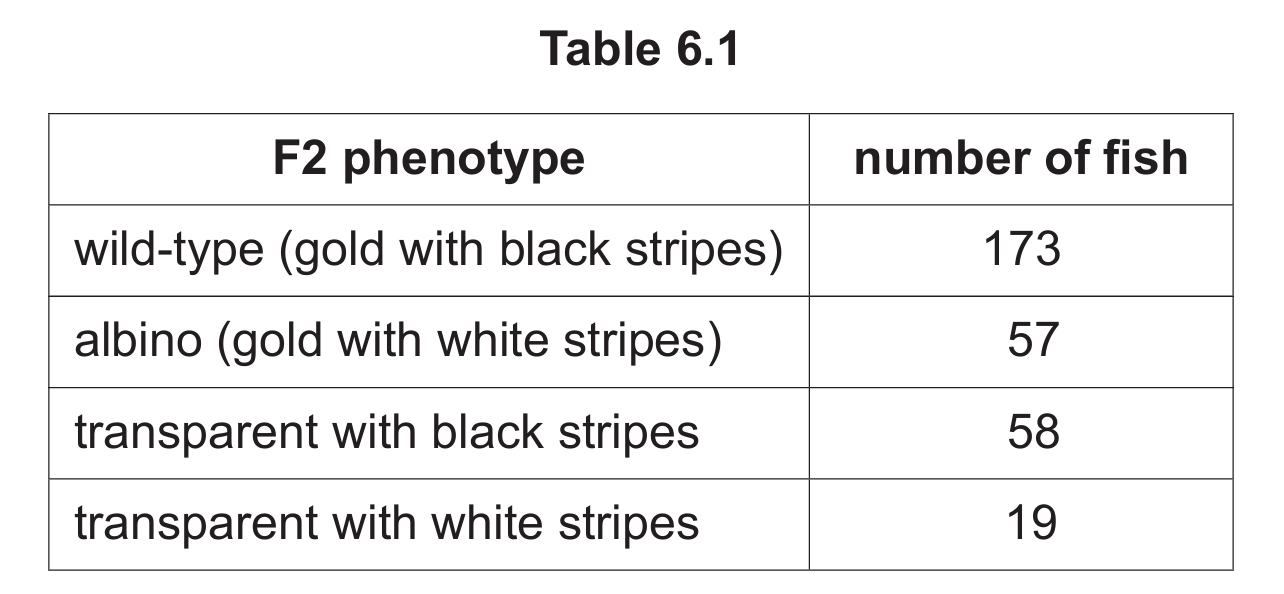
(a) (i) The fish breeder concluded that the new transparent phenotype had occurred because of a mutation.
Explain how the results in Table 6.1 support this conclusion.
(ii) State the approximate whole-number ratio shown by the results in Table 6.1.
(iii) Explain what the results in Table 6.1 show about the genes and alleles that determine the wild-type, albino and the two different transparent phenotypes in tiger barbs.
(b) The F1 tiger barbs all looked the same but the F2 offspring showed variation. The F2 offspring showed four different phenotypes.
Describe the processes that occurred during meiosis in the F1 fish that allowed this variation to occur.
▶️ Answer/Explanation
(a) (i)
The results support the mutation conclusion because:
- The transparent phenotype reappears in the F2 generation after being absent in the F1 generation, showing it is inherited rather than being an environmental effect.
- The pattern of inheritance follows Mendelian genetics, suggesting the transparent trait is controlled by a genetic factor (a new allele) rather than being a random environmental change.
- The appearance of a new phenotype (transparent) that wasn’t present in either original parent suggests a genetic change occurred.
(a) (ii)
The approximate ratio is 9:3:3:1 (wild-type : albino : transparent with black stripes : transparent with white stripes).
(a) (iii)
The results show that:
- There are two genes involved in determining these phenotypes.
- The genes are on separate chromosomes (not linked) as they show independent assortment.
- Each gene has two alleles: one dominant and one recessive.
- For the color pattern:
- Wild-type (gold with black stripes) is dominant for both genes.
- Albino (gold with white stripes) is recessive for the stripe color gene but dominant for the body color gene.
- Transparent with black stripes is recessive for the body color gene but dominant for the stripe color gene.
- Transparent with white stripes is recessive for both genes.
- The genes are autosomal (not sex-linked) as the ratios appear in both sexes.
(b)
The variation in F2 occurs because of these meiotic processes:
- Independent assortment of chromosomes during metaphase I of meiosis, where homologous chromosome pairs align randomly at the cell equator.
- Crossing over during prophase I, where homologous chromosomes exchange genetic material, creating new combinations of alleles.
- Random segregation of homologous chromosomes during anaphase I, where each pair separates independently of other pairs.
- Separation of sister chromatids in meiosis II, producing four genetically different gametes from each parent.
- When these gametes combine randomly during fertilization, they produce the observed 9:3:3:1 phenotypic ratio in the F2 generation.
This genetic recombination during meiosis explains why the F1 fish (all heterozygous) looked uniform, while their offspring showed variation through new combinations of the parental alleles.
Topic: 15.1
When an impulse arrives at a neuromuscular junction, it stimulates a muscle fibre of striated muscle to contract.
(a) (i) Outline the similarities in structure between a neuromuscular junction and a cholinergic synapse.
(ii) The hydrolysis of ATP during muscle contraction releases inorganic phosphate (Pi). Calcium ions (Ca2+) can combine with Pi in the sarcoplasmic reticulum to form insoluble calcium phosphate. This may result in fewer power strokes occurring in sarcomeres.
Suggest why calcium phosphate formation in the sarcoplasmic reticulum may result in fewer power strokes occurring in sarcomeres.
(b) Adrenaline is a hormone that can affect muscle contraction. Adrenaline binds to G-protein-coupled receptors on T-tubule membranes.
The cell signalling pathway that occurs in response to the binding of adrenaline is similar to the pathway that occurs in liver cells in response to the binding of glucagon.
Fig. 7.1 is an outline of the cell signalling pathway of adrenaline.
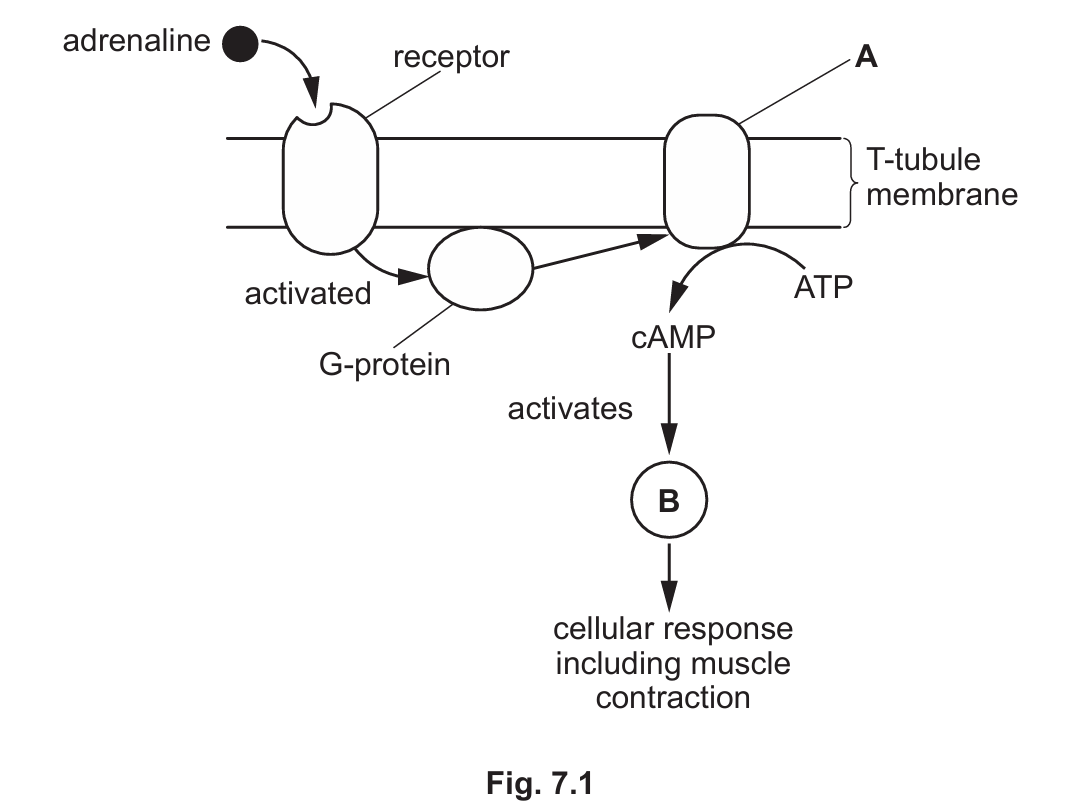
Identify the molecules represented by A and B in Fig. 7.1.
▶️ Answer/Explanation
(a)(i) Similarities between neuromuscular junction and cholinergic synapse:
1. Both contain vesicles of acetylcholine which are released into the synaptic cleft.
2. Both have numerous mitochondria to provide ATP for neurotransmitter release.
3. Both have distinct presynaptic and postsynaptic membranes separated by a synaptic cleft.
4. Both have receptors for acetylcholine on the postsynaptic membrane.
5. Both involve calcium ion entry into the presynaptic terminal to trigger neurotransmitter release.
6. Both involve sodium ion channels opening in the postsynaptic membrane during depolarization.
Explanation: The neuromuscular junction is essentially a specialized type of cholinergic synapse between a motor neuron and muscle fiber. They share the same basic structure and mechanism of signal transmission using acetylcholine as the neurotransmitter.
(a)(ii) Why calcium phosphate formation reduces power strokes:
1. Formation of calcium phosphate reduces the number of free calcium ions available in the sarcoplasmic reticulum.
2. Fewer calcium ions are released into the sarcoplasm during muscle stimulation.
3. Reduced calcium means less binding to troponin molecules on the actin filaments.
4. Without sufficient calcium-troponin binding, tropomyosin doesn’t move enough to expose myosin binding sites on actin.
5. Fewer actin-myosin cross bridges can form, resulting in fewer power strokes.
6. Additionally, the trapped phosphate ions are unavailable for ATP regeneration, further limiting muscle contraction.
Explanation: Calcium ions are crucial for initiating the contraction cycle by binding to troponin. When they become sequestered as calcium phosphate, this essential trigger is diminished, directly reducing the number of power strokes that can occur in the sarcomeres.
(b) Identification of molecules A and B:
A: Adenylyl cyclase
B: Protein kinase A
Explanation: In the adrenaline signaling pathway shown in Fig. 7.1, the activated G-protein stimulates adenylyl cyclase (A) to convert ATP to cAMP. The cAMP then activates protein kinase A (B), which phosphorylates various target proteins to produce the cellular response, including enhanced muscle contraction. This is the same basic pathway used by glucagon in liver cells.
Topic: 13.1
Environmental conditions such as light intensity affect plant physiology.
(a) Use the letter X to identify a point on the sketch graph in Fig. 8.1 where light intensity is acting as a limiting factor on the rate of photosynthesis.
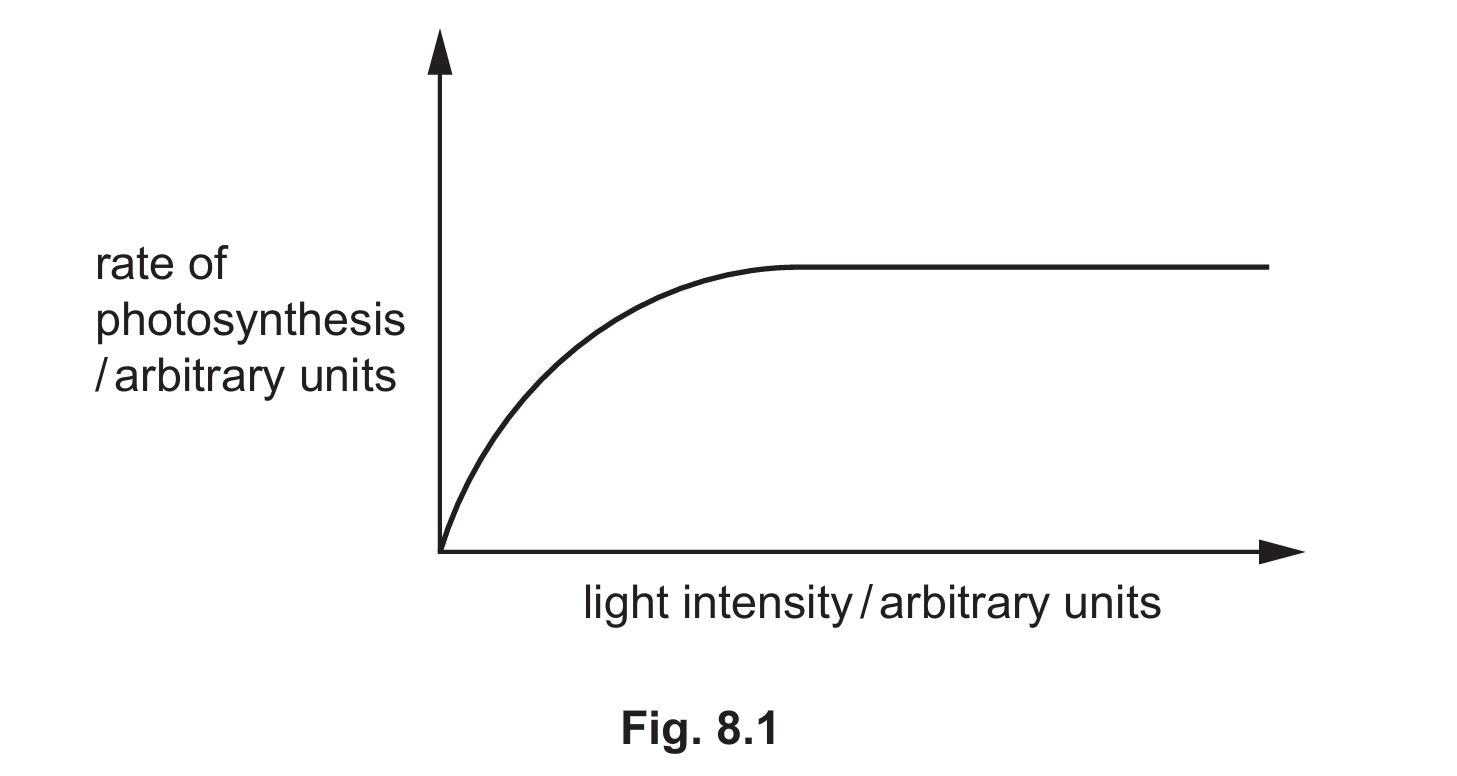
(b) An experiment investigated how light intensity affected gene expression in kale, Brassica oleracea sabellica.
- Two groups of kale plants were grown, with one group in high light intensity and one group in low light intensity. All other conditions were standardized.
- After the same period of time, messenger RNA (mRNA) was extracted from each group of kale plants.
- The mRNA was used to produce cDNA.
- The cDNA was hybridized with probes for 89,621 kale genes on a microarray.
- The microarrays showed which genes were switched on (expressed) in each set of conditions.
The results showed that expression of 18% of the genes was affected by light intensity.
- 14% of the genes were switched on only in high light intensity.
- 4% were switched on only in low light intensity.
(i) State the name of the enzyme that produces cDNA from an mRNA template.
(ii) State the name of the type of proteins that control gene expression in plants.
(iii) Suggest why more genes were switched on only in plants growing in high light intensity compared to fewer genes that were switched on only in plants growing in low light intensity.
▶️ Answer/Explanation
(a) Letter X should be placed on the rising slope of the graph, to the left of the point where the line plateaus.
Explanation: Light intensity acts as a limiting factor when increasing it leads to an increase in the rate of photosynthesis (the rising part of the curve). Once the line plateaus, other factors like CO2 concentration or temperature become limiting instead.
(b)(i) Reverse transcriptase
Explanation: This enzyme synthesizes complementary DNA (cDNA) from an mRNA template in a process called reverse transcription. It’s essential for converting the mRNA extracted from the kale plants into stable cDNA that can be used for microarray analysis.
(b)(ii) Transcription factors
Explanation: These are proteins that bind to specific DNA sequences to control the rate of transcription of genetic information from DNA to mRNA. They can either activate or repress gene expression in response to environmental conditions like light intensity.
(b)(iii) There are several possible reasons:
Explanation:
- In high light intensity, plants can photosynthesize more and thus need to upregulate more genes for photosynthetic enzymes (like Rubisco) and proteins involved in the light-dependent reactions.
- High light may require additional protective pigments (like carotenoids) and enzymes to prevent photodamage, hence more genes are activated.
- With more energy available, plants can activate genes for growth and development, including those for making sucrose, starch, and amino acids.
- Low light conditions might primarily require genes for light-harvesting complexes and shade-avoidance responses, which are fewer in number compared to the diverse processes needed in high light.
- The greater energy availability in high light allows for expression of a wider range of genes involved in various metabolic pathways beyond just basic survival.
This differential gene expression helps plants optimize their growth and survival under different light conditions.
Topic: 18.3
Green lacewings are a family of insects with more than 1300 species.
The common green lacewing, Chrysoperla carnea, is shown in Fig. 9.1.

(a) Green lacewings have sense organs, known as tympanal organs, that detect sound.
The tympanal organ of green lacewings has evolved to detect the high frequency sounds that bats make when they are hunting. Bats eat green lacewings.
When a green lacewing senses the presence of a bat, it moves away or closes its wings in flight to escape.
(i) Outline how the tympanal organ of green lacewings could have evolved by natural selection.
(ii) When high frequency sound is detected, the receptor cells in the tympanal organ stimulate the transmission of impulses in sensory neurones.
Describe the sequence of events that results in an action potential in a sensory neurone.
The first event in the sequence has been given for you.
Calcium ions enter the cytoplasm of the receptor cell
(b) Two species of green lacewing, C. carnea and C. downesi, evolved from a common ancestor.
The two species have populations with overlapping distributions in parts of North America.
Table 9.1 shows a comparison of the characteristics of overlapping populations of the two species.
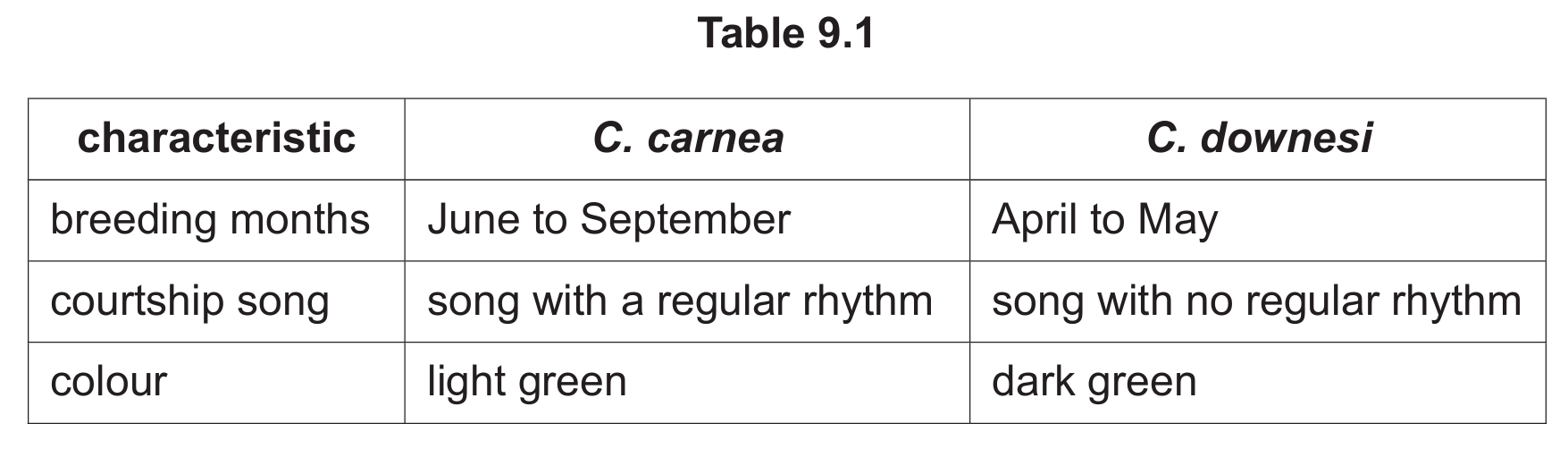
Suggest how speciation occurred to produce the two different species of green lacewing.
▶️ Answer/Explanation
(a)(i) Outline how the tympanal organ of green lacewings could have evolved by natural selection:
1. Initially, random mutations occurred in some lacewings that gave them limited ability to detect high frequency sounds.
2. The selection pressure came from bat predation – those lacewings that could detect bats’ high frequency hunting calls had a survival advantage.
3. These surviving lacewings with better hearing were more likely to reproduce and pass on their advantageous alleles to offspring.
4. Over many generations, the tympanal organs became more sensitive to high frequencies through the accumulation of beneficial mutations.
5. Eventually, the entire population evolved this adaptation, making them better at avoiding bat predation.
(a)(ii) Sequence of events resulting in an action potential:
1. Calcium ions enter the cytoplasm of the receptor cell (as given in the question).
2. This causes vesicles containing neurotransmitter to move toward and fuse with the presynaptic membrane.
3. Neurotransmitter (likely glutamate or acetylcholine) is released into the synaptic cleft via exocytosis.
4. The neurotransmitter binds to ligand-gated sodium channels on the sensory neurone’s membrane.
5. Sodium ions (Na+) enter the sensory neurone, causing depolarization of the membrane.
6. If the depolarization reaches threshold potential (-55mV), voltage-gated sodium channels open, triggering an action potential.
7. The action potential propagates along the sensory neurone via saltatory conduction (in myelinated neurones) or continuous conduction (in unmyelinated neurones).
(b) Speciation in green lacewings:
This appears to be an example of sympatric speciation, where the two species evolved while living in the same geographic area. The process likely involved:
1. The ancestral population developed genetic variations through mutations, including differences in breeding seasons, courtship songs, and coloration.
2. Some individuals began breeding in spring (April-May) while others bred in summer (June-September), creating temporal reproductive isolation.
3. The different breeding seasons led to the evolution of distinct courtship songs – regular rhythm for summer breeders and irregular for spring breeders.
4. The coloration differences (light vs dark green) may have provided camouflage in slightly different habitats or at different times of year.
5. Over time, these behavioral and physical differences became so pronounced that the two groups could no longer interbreed, even when living in the same areas.
6. The lack of gene flow between the groups allowed them to diverge genetically, eventually forming distinct species.
This speciation mechanism is supported by the overlapping distributions and the clear differences in mating behaviors that would prevent interbreeding.
Topic: 19.3
Medicine is defined as the diagnosis, treatment and prevention of disease.
Outline the different ways in which genetic technology can be applied to medicine, with reference to named diseases.
▶️ Answer/Explanation
Genetic technology has revolutionized medicine in several ways:
1. Genetic Engineering/Recombinant DNA Technology:
This involves inserting human genes into microorganisms to produce therapeutic proteins. For example:
- Insulin production for diabetes treatment – Human insulin gene is inserted into bacteria or yeast, which then produce human insulin.
- Factor VIII for hemophilia treatment – The clotting factor protein is produced by genetically modified organisms.
- Adenosine deaminase (ADA) for Severe Combined Immunodeficiency (SCID) – This enzyme is crucial for immune system function.
2. Genetic Screening/Diagnosis:
DNA testing allows early detection of genetic disorders:
- BRCA1/BRCA2 genes testing for breast cancer risk assessment.
- Detection of Huntington’s disease through analysis of CAG repeats in the HTT gene.
- Identification of cystic fibrosis through CFTR gene mutation analysis.
Positive diagnoses can lead to early interventions, lifestyle changes, or preventive measures.
3. Gene Therapy:
This involves introducing normal genes into patients’ cells to compensate for defective ones:
- Treatment of Leber’s congenital amaurosis (LCA), an inherited eye disease, by delivering normal RPE65 gene to retinal cells.
- Treatment of SCID by introducing functional ADA gene into patients’ bone marrow cells.
4. Other Applications:
- Xenotransplantation: Using genetically modified pigs as potential organ donors for humans.
- Recombinant vaccines: Like the hepatitis B vaccine produced by yeast cells.
- mRNA vaccines: Such as those developed for COVID-19, which represent a new era in vaccine technology.
- Monoclonal antibody production: Using recombinant DNA technology to produce targeted therapies for cancer and autoimmune diseases.
These applications demonstrate how genetic technology has transformed modern medicine, enabling more precise diagnoses, effective treatments, and preventive strategies for numerous diseases.
