GEN 3.4 Mutations- Pre AP Biology Study Notes - New Syllabus.
GEN 3.4 Mutations- Pre AP Biology Study Notes
GEN 3.4 Mutations- Pre AP Biology Study Notes – New Syllabus.
LEARNING OBJECTIVE
GEN 3.4(a) Describe how changes in DNA sequences may affect protein structure and function.
GEN 3.4(b) Create and/or use models to explain the consequences of changes in DNA.
GEN 3.4(c) Analyze data to make predictions about how changes in DNA affect an organism’s phenotype.
Key Concepts:
GEN 3.4.1 Mutations are heritable changes to DNA sequences.
a. Mutations are random changes in DNA sequences that may occur as a result of errors during replication or the effects of environmental mutagens (e.g., UV light, X-rays, and carcinogens).
b. A change in a DNA sequence occurs when a nucleotide is substituted into the original sequence (causing a point mutation) or inserted into or deleted from the sequence (causing a frameshift mutation).
c. Depending on how the changes impact gene expression, mutations may cause negative disruption in gene and protein function, have little to no effect on organisms, or produce beneficial variation.
How Changes in DNA Sequences May Affect Protein Structure and Function
🌿 Introduction
DNA stores genetic information in the sequence of its nitrogen bases:
A, T, G, C
The specific order of these bases determines the amino acid sequence of a protein.
If the DNA sequence changes, the information may change.
A permanent change in the DNA sequence is called a mutation.
Mutations can affect:
- Protein structure
- Protein function
- Cell function
- Organism phenotype
🧠 DNA Sequence Determines Protein Structure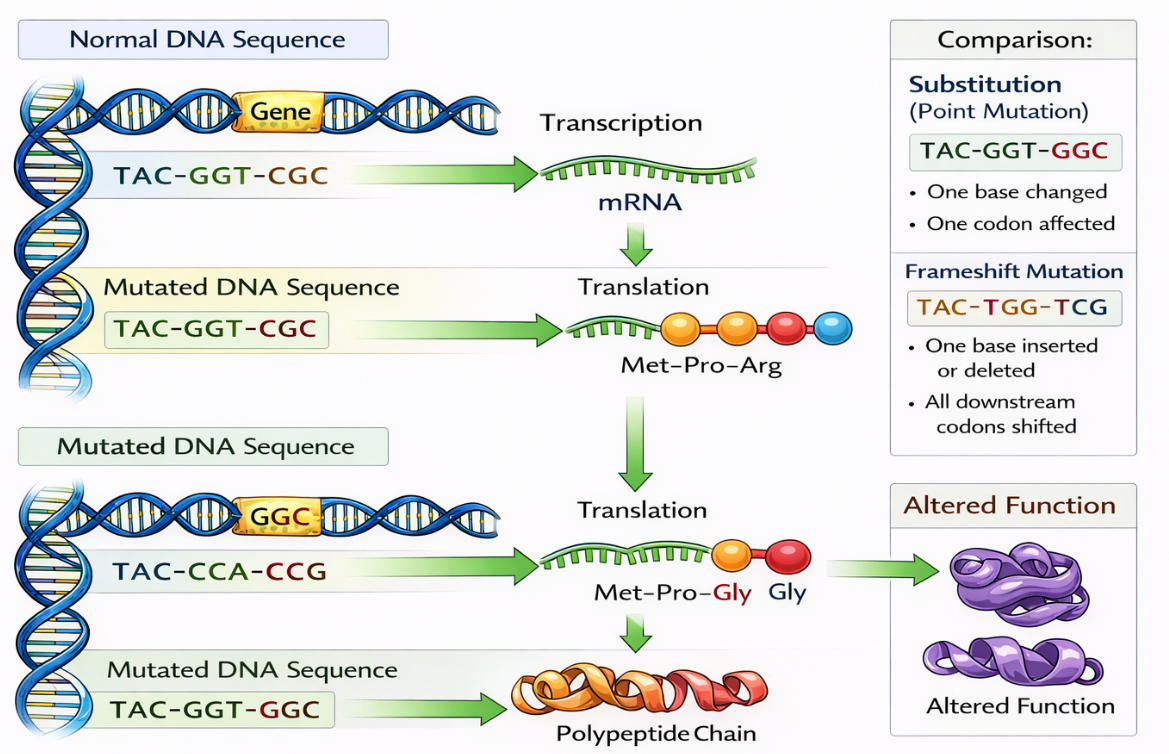
The sequence of bases in DNA determines:
DNA → mRNA → Amino acid sequence → Protein structure
Step-by-step:
- DNA is transcribed into mRNA.
- mRNA is read in codons (three bases at a time).
- Each codon specifies one amino acid.
- Amino acids are linked to form a polypeptide.
- The polypeptide folds into a specific 3D structure.
The final shape of the protein determines its function.
So, the base sequence in DNA ultimately determines how a protein works.
🧬 What Happens When DNA Changes?
If a nucleotide in DNA changes, the codon in mRNA may change.
This may cause:
- A different amino acid to be added
- No change in amino acid
- A stop codon to appear early
Any of these outcomes can affect the protein.
🧪 Types of DNA Sequence Changes
Substitution (Point Mutation)
One nucleotide is replaced by another.
Example:
Original DNA: A T G
Mutated DNA: A T A
Only one base changed.
Possible effects:
- May change one amino acid
- May not change amino acid at all
- May create a stop codon
Because only one codon is affected, impact is often limited.
Insertion
An extra nucleotide is added.
This changes the reading frame of the codons.
Example:
Original: A T G – C C A
After insertion: A T G – T C C – A
Now all codons after the insertion are different.
This is called a frameshift mutation.
Deletion
A nucleotide is removed.
This also shifts the reading frame.
Frameshift mutations often:
- Change many amino acids
- Create premature stop codons
- Produce nonfunctional proteins
They are usually more disruptive than substitutions.
🧠 How Amino Acid Changes Affect Protein Structure
Proteins depend on their amino acid sequence for correct folding.
Each amino acid has unique chemical properties:
- Size
- Charge
- Polarity
If a mutation changes one amino acid:
- The folding pattern may change
- Chemical interactions may change
- The shape may change
Since protein function depends on shape, function may change.
🧬 Effects on Protein Function
Disrupted (Negative Effect)
- Enzyme active site altered
- Protein cannot bind substrate
- Protein may not fold correctly
- Loss of function
This can lead to disorders.
No Effect (Neutral Mutation)
- Amino acid may not change
- Or similar amino acid substituted
- Protein function remains unchanged
Because of redundancy in genetic code, some mutations are silent.
Beneficial Effect
- Protein works more efficiently
- New function develops
- May improve survival
Beneficial mutations contribute to variation in populations.
🧠 Overall Cause-and-Effect Chain
Mutation in DNA → Change in mRNA codon → Change in amino acid → Change in protein folding → Change in protein function → Change in phenotype
This is how DNA sequence changes can affect organism traits.
📊 Summary Table
| DNA Change | Protein Impact | Possible Outcome |
|---|---|---|
| Substitution | One amino acid may change | Mild to moderate effect |
| Insertion | Frameshift | Often severe |
| Deletion | Frameshift | Often severe |
| Silent change | No amino acid change | No effect |
| Stop codon creation | Shortened protein | Likely nonfunctional |
📦 Quick Recap
Mutation = change in DNA sequence
DNA change → codon change → amino acid change
Amino acid sequence determines protein structure
Structure determines function
Effects may be negative, neutral, or beneficial
Frameshift mutations often more severe
Using Models to Explain the Consequences of Changes in DNA
🌿 Introduction
A mutation is a change in the DNA sequence.
But to truly understand its impact, we must use a model that traces the change from:
DNA → mRNA → Amino acid sequence → Protein → Phenotype
A mutation model helps us visualize how even a small DNA change can influence the final trait.
The goal of this objective is not just to identify mutations, but to demonstrate their consequences clearly using a step-wise model.
🧠 What a Mutation Model Must Include
A complete model explaining consequences of DNA change must show:
- Original DNA sequence
- Mutated DNA sequence
- Resulting mRNA sequences
- Codon changes
- Amino acid changes
- Resulting protein differences
- Possible phenotypic outcome
The model must connect molecular change to observable effect.
🧬 Model 1 – Substitution (Point Mutation)
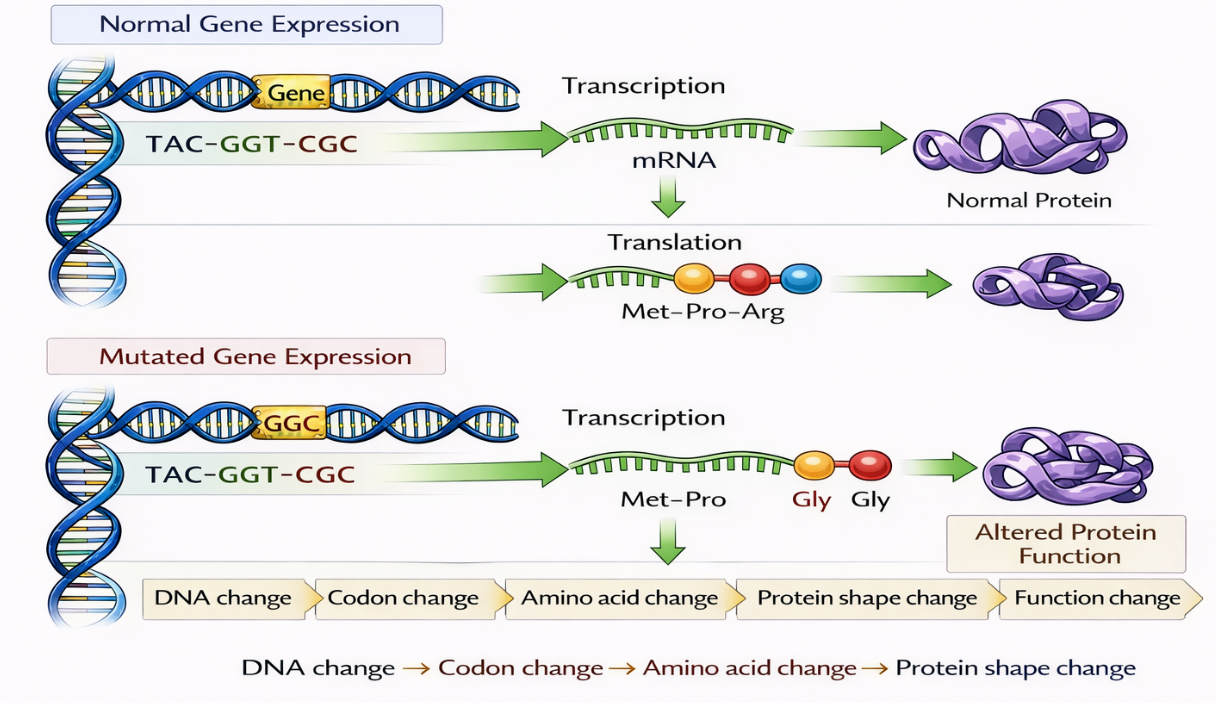
Step 1 – Original DNA
Example:
DNA: T A C – G A A – C T T
Transcription produces:
mRNA: A U G – C U U – G A A
Translation produces:
Met – Leu – Glu
Protein forms normally.
Step 2 – Substituted DNA
Now change one base:
DNA: T A C – G A T – C T T
mRNA becomes:
A U G – C U A – G A A
One codon changes.
Now amino acid sequence changes at one position.
Possible consequences:
- One amino acid replaced
- Protein structure slightly altered
- Function may remain same or slightly altered
This model shows:
Small DNA change → Small protein change.
🧬 Model 2 – Frameshift Mutation (Insertion or Deletion)
Step 1 – Original DNA
DNA: T A C – G A A – C T T
mRNA: A U G – C U U – G A A
Amino acids: Met – Leu – Glu
Step 2 – Insertion of One Base
DNA: T A C – G A A – C T T
Insert “A” after first codon: T A C – A G A – A C T – T
Now mRNA reading frame shifts.
All codons after insertion change.
Amino acid sequence changes completely after mutation.
Possible consequences:
- Many incorrect amino acids
- Premature stop codon
- Shortened protein
- Nonfunctional protein
This model demonstrates:
One base insertion → Entire downstream sequence affected.
Frameshift mutations often have more severe consequences.
🧠 Demonstrating Protein-Level Consequences
A mutation model must show protein effects clearly.
Possible outcomes:
Loss of Function
- Protein cannot fold properly
- Active site altered
- Enzyme stops working
- Trait may be negatively affected
No Significant Effect
- Amino acid not changed
- Similar amino acid substituted
- Protein function unchanged
Some mutations are silent.
Altered or Beneficial Function
- Slight structural change improves efficiency
- New functional variation arises
Such mutations contribute to genetic diversity.
🧬 Connecting Model to Phenotype
Mutation → Altered protein → Changed cellular function → Changed phenotype
Example:
Mutation in pigment-producing enzyme → Enzyme inactive → No pigment formed → Different coloration phenotype
This demonstrates how molecular change leads to observable change.
🧠 How to Use the Model to Predict Outcomes
When given DNA data:
Step 1
Identify the mutation type.
Step 2
Determine if reading frame changes.
Step 3
Translate new codons.
Step 4
Compare amino acid sequences.
Step 5
Predict effect on protein structure.
Step 6
Predict phenotypic consequence.
This structured modeling approach allows prediction of biological impact.
📊 Comparison Table of Mutation Models
| Mutation Type | DNA Change | Protein Impact | Likely Consequence |
|---|---|---|---|
| Substitution | One base replaced | One amino acid affected | Mild to moderate |
| Insertion | Extra base added | Frameshift | Often severe |
| Deletion | Base removed | Frameshift | Often severe |
| Silent | Codon unchanged in meaning | No amino acid change | No effect |
🧠 Big Concept
A mutation model demonstrates:
DNA sequence change → mRNA codon change → Amino acid change → Protein structure change → Functional consequence → Phenotype impact
The model connects molecular biology to observable traits.
📦 Quick Recap
Substitution affects one codon
Frameshift affects many codons
Protein structure determines function
Function determines phenotype
Effects may be harmful, neutral, or beneficial
Analyze Data to Predict How DNA Changes Affect Phenotype
🌿 Introduction
DNA stores genetic information in the sequence of bases.
When the DNA sequence changes, the protein encoded by that gene may change.
Since proteins determine traits, changes in DNA can affect phenotype.
This objective focuses on:
- Analyzing sequence data
- Identifying mutation type
- Predicting protein impact
- Predicting phenotypic outcome
🧠 Step-by-Step Method to Analyze Mutation Data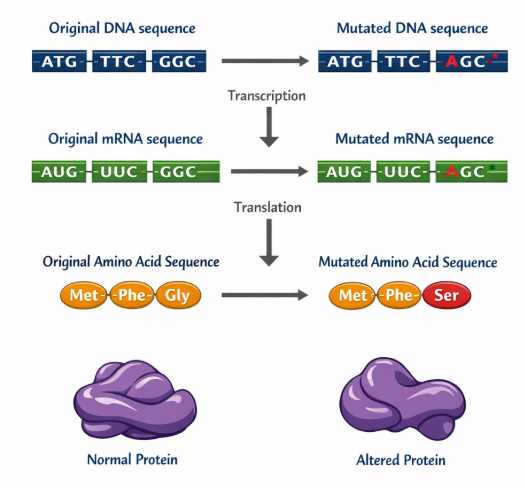
When given DNA data, always follow this structured process:
Step 1 – Compare Original and Mutated DNA
Look for:
- Substitution
- Insertion
- Deletion
Identify exactly where the change occurs.
Step 2 – Determine the Mutation Type
One base replaced → Substitution
One base added → Insertion
One base removed → Deletion
If insertion or deletion occurs, check if reading frame shifts.
Frameshift mutations usually have larger effects.
Step 3 – Transcribe to mRNA
Use base pairing rules:
DNA A → RNA U
DNA T → RNA A
DNA G → RNA C
DNA C → RNA G
Write both original and mutated mRNA sequences.
Step 4 – Translate mRNA into Amino Acids
Group mRNA into codons (3 bases).
Determine amino acid sequence for:
- Original
- Mutated
Compare the two sequences.
Step 5 – Predict Protein Impact
Ask:
- Did one amino acid change?
- Did multiple amino acids change?
- Did a stop codon appear early?
Possible outcomes:
- No change in amino acid → likely no effect
- One amino acid change → possible minor effect
- Frameshift → likely major disruption
- Early stop codon → shortened protein
Step 6 – Predict Phenotypic Effect
Now connect to phenotype.
Protein function determines trait.
If protein:
- Functions normally → phenotype unchanged
- Partially altered → phenotype slightly altered
- Nonfunctional → phenotype significantly affected
🧬 Example Data Analysis
Example 1 – Substitution with No Amino Acid Change
Original DNA: T A C – G A A – C T T
Mutated DNA: T A C – G A G – C T T
After transcription and translation: Amino acid sequence remains same.
Prediction: Protein unchanged → Phenotype unchanged.
This is a neutral mutation.
Example 2 – Substitution Changing Amino Acid
Original DNA: T A C – G A A – C T T
Mutated DNA: T A C – G A T – C T T
One amino acid changes.
Prediction: Protein structure may slightly change → Function may be altered → Phenotype may change slightly.
Example 3 – Frameshift Mutation
Original DNA: T A C – G A A – C T T
Insert one base: T A C – A G A – A C T – T
All codons after mutation change.
Prediction: Many amino acids change → Protein likely nonfunctional → Phenotype significantly altered.
🧠 Predicting Severity of Mutation
Severity depends on:
- Location of mutation
- Type of mutation
- Whether protein is essential
- Whether mutation affects active site
Frameshift mutations are generally more severe.
Silent substitutions often have no effect.
🧬 Connecting Genotype to Phenotype
Genotype = DNA sequence
Phenotype = observable trait
Mutation changes genotype.
If genotype change alters protein structure, phenotype may change.
So, prediction always follows:
DNA change → Protein change → Trait change
📊 Summary Table
| Mutation Type | Protein Impact | Predicted Phenotype |
|---|---|---|
| Silent substitution | No amino acid change | No effect |
| Missense substitution | One amino acid change | Possible minor effect |
| Nonsense mutation | Early stop codon | Likely severe |
| Frameshift | Many amino acids altered | Often severe |
📦 Quick Recap
Identify mutation type
Transcribe to mRNA
Translate to amino acids
Determine protein impact
Predict phenotype change
