Pre AP Biology -EVO 1.2 Classifying Evolutionary Relationships - MCQ Exam Style Questions -New Syllabus 2025-2026
Pre AP Biology -EVO 1.2 Classifying Evolutionary Relationships – MCQ Exam Style Questions – New Syllabus 2025-2026
Pre AP Biology -EVO 1.2 Classifying Evolutionary Relationships – MCQ Exam Style Questions – Pre AP Biology – per latest Pre AP Biology Syllabus.
Question
b. Arthropoda
c. Echinodermata
d. Porifera
▶️ Answer/Explanation
The correct answer is a. Cnidaria.
A medusa is one of the two primary body forms found in the phylum Cnidaria.
It is characterized by a bell-shaped, free-swimming body with tentacles hanging downward.
Common examples of the medusa form include jellyfish.
The other primary body form in this phylum is the sessile polyp, such as sea anemones.
Phyla like Arthropoda, Echinodermata, and Porifera do not exhibit the medusa body plan.
Question
▶️ Answer/Explanation
The correct answer is d. $4$.
Basal phyla are those that diverged early from the main animal lineage.
Scientists generally identify $4$ distinct basal animal phyla.
These include Porifera (sponges) and Cnidaria (jellyfish and corals).
They also include Ctenophora (comb jellies) and Placozoa.
These groups lack the complex bilateral symmetry found in Bilateria.
They represent the most primitive branches of the animal phylogenetic tree.
Question
▶️ Answer/Explanation
The correct option is b.
Key innovations refer to traits that evolved within the animal lineage to drive diversification.
Heterotrophy and multicellularity are ancestral traits shared with fungi or protist ancestors.
The evolution of symmetry (radial and bilateral) allowed for directed movement and cephalization.
The development of specialized tissues enabled complex organ systems and physiological division of labor.
These two features distinguishes Parazoa (sponges) from Eumetazoa (true animals).
Therefore, symmetry and tissue differentiation are the primary evolutionary “milestones” listed.
Question
▶️ Answer/Explanation
The correct answer is a. Opisthokonts.
Opisthokonta is a broad group of eukaryotes that includes both the Animal and Fungus kingdoms.
This supergroup is characterized by cells that possess a single posterior flagellum at some point in their life cycle.
Animals are specifically closely related to choanoflagellates, which are protists within this supergroup.
Other groups like Rhizaria or Stramenopiles belong to different eukaryotic lineages (such as the SAR clade).
Molecular evidence and structural similarities confirm that animals share a common ancestor within the Opisthokont lineage.
Question
▶️ Answer/Explanation
The correct answer is d. monophyletic taxa or lineages only.
In modern cladistics, a valid biological classification must reflect evolutionary history.
To achieve this, scientists identify monophyletic groups, also known as clades.
A monophyletic group includes a common ancestor and all of its descendants.
Paraphyletic and polyphyletic groups are avoided as they do not represent complete evolutionary lineages.
By naming only monophyletic groups, the classification becomes a direct map of the phylogenetic tree.
Question
▶️ Answer/Explanation
The correct option is d.
Maximum parsimony is a character-based method that seeks the simplest explanation for observed data.
It involves grouping organisms based on shared derived characters (synapomorphies) rather than ancestral ones.
The “parsimony assumption” specifically dictates choosing the tree topology that requires the minimum total number of character state transitions.
In molecular terms, this means selecting the tree that minimizes the number of base substitutions or mutations.
By reducing the number of hypothesized evolutionary changes, researchers avoid unnecessary complexity in the phylogenetic model.
This approach assumes that the most likely evolutionary path is the one involving the fewest independent genetic events.
Question
b. derived homoplasious
c. ancestral homologous
d. ancestral homoplasious
▶️ Answer/Explanation
The correct answer is a. derived homologous.
Cladistics relies on synapomorphies, which are shared derived characters.
Homologous traits indicate a shared evolutionary ancestry between groups.
Derived traits are those that evolved in the lineage leading to a specific clade.
Ancestral traits (plesiomorphies) are rejected because they do not distinguish sub-groups.
Homoplasious traits are ignored as they result from convergent evolution, not ancestry.
Systematists use these shared derived traits to define monophyletic groups.
Question
▶️ Answer/Explanation
The correct option is d.
Molecular clocks rely on the accumulation of mutations over time to estimate evolutionary divergence.
A reliable clock requires a reasonably constant rate of mutation to act as a consistent “ticking” mechanism.
Noncoding regions of DNA are often preferred because they are not subject to intense natural selection.
In these regions, neutral mutations can accumulate without affecting the organism’s fitness.
This allows the number of genetic differences to be proportional to the time since two species shared a common ancestor.
Coding regions and proteins are less ideal as selection pressures can cause sudden changes in mutation rates.
Question
b. its evolutionary history
c. its domain
d. its distribution
▶️ Answer/Explanation
The correct option is b. its evolutionary history.
A phylogenetic tree is a diagram that represents evolutionary relationships among organisms.
It reflects how different species evolved from common ancestors over time.
While classification (a) is related, it is the naming system based on these relationships.
The domain (c) is merely the broadest category of biological classification.
Distribution (d) refers to geographic location, not ancestral lineage.
Therefore, the tree specifically maps out the phylogeny or historical descent of the group.
Question
▶️ Answer/Explanation
The correct answer is c. molecular phylogenetic analyses.
Scientists compared the nucleotide sequences of $\text{HIV-1}$ and $\text{HIV-2}$ with Simian Immunodeficiency Viruses ($\text{SIV}$).
By constructing evolutionary trees, they found $\text{HIV-1}$ is more closely related to $\text{SIV}_{\text{cpz}}$ from chimpanzees.
$\text{HIV-2}$ was found to be more closely related to $\text{SIV}_{\text{sm}}$ from sooty mangabeys.
These trees showed the viruses entered humans multiple times from non-human primates.
Therefore, the analysis proved the virus did not evolve its distinct strands within human hosts.
This specific method of using molecular data to determine evolutionary history is known as molecular phylogenetics.
Question
▶️ Answer/Explanation
The correct answer is b. systematics.
Systematics allowed researchers to distinguish between identical-looking mosquito species.
It was discovered that only specific members of the Anopheles maculipennis complex transmitted malaria.
Some species lived in brackish water, while others lived in fresh water or fed on livestock instead of humans.
By identifying these specific vectors, control efforts were accurately targeted rather than wasted.
This precise biological classification was essential for the successful eradication of the disease in Europe.
Question
▶️ Answer/Explanation
The correct option is (a).
Bacteria exhibit a vast diversity in genome size, ranging from approximately $1.6 \times 10^5$ to over $1.0 \times 10^7$ base pairs.
Option (b) is incorrect as many plants and amphibians have genomes much larger than the $3.2 \times 10^9$ base pairs found in humans.
Option (c) is false due to the C-value paradox, where genome size does not correlate with biological complexity.
Option (d) is incorrect because large eukaryotic genomes often consist of non-coding repetitive DNA rather than a higher count of functional genes.
Therefore, the wide variation in bacterial genome sizes is the most accurate biological description among the choices.
Questions
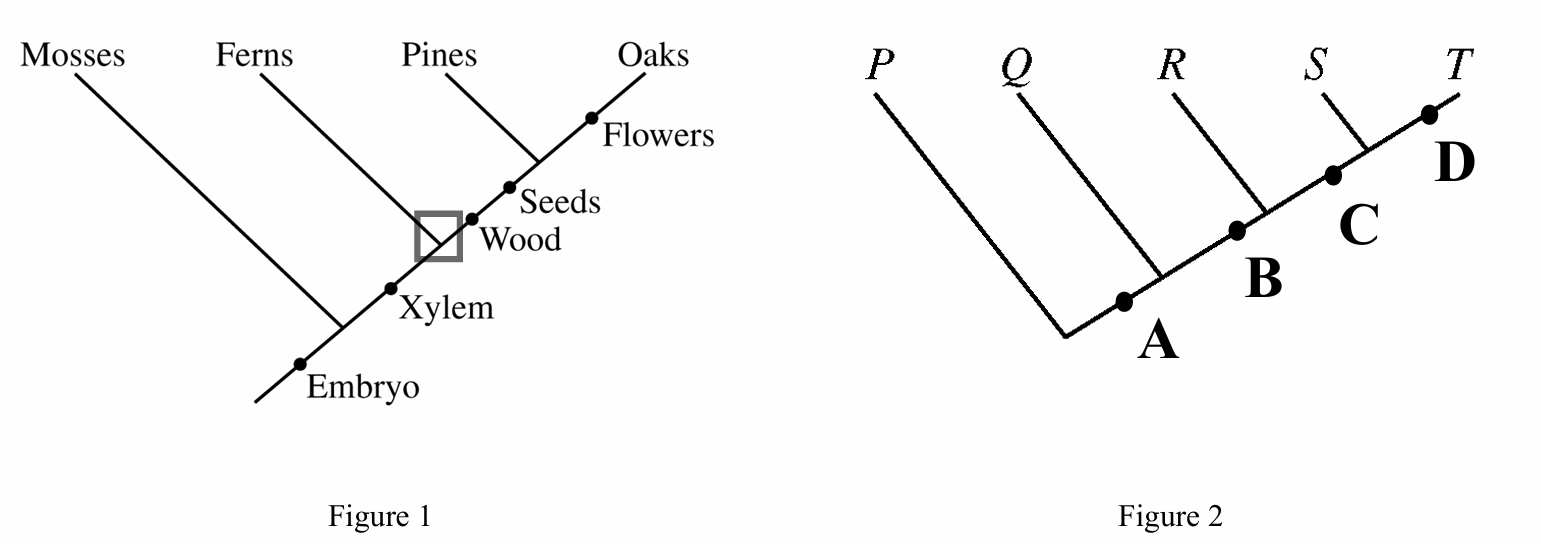
B. Seeds and wood only
C. Embryo and xylem only
D. Embryo, xylem, wood, and seeds
B. Ferns
C. Pines
D. Oaks
B. Ferns and Pines
C. Pines and Oaks
D. Oaks and Mosses
B. Species \(P\) and \(T\) do not share a common ancestor.
C. Species \(S\) evolved from species \(R\).
D. Species \(S\) is more closely related to species \(T\) than to species \(R\).
B. Characteristic B is found in species \(Q\), \(R\), \(S\), and \(T\), but not in species \(P\).
C. Characteristic D is the most ancestral trait.
D. Characteristic C is more derived than characteristic B.
▶️ Answer/Explanation
The cladogram shows the evolutionary order of traits relative to the branching of species.
The trait Xylem appears on the main line after the Mosses branch off but before the Ferns branch off.
This means Mosses do not possess Xylem (ancestral state), while Ferns and all subsequent lineages (Pines, Oaks) do possess Xylem.
Traits like Wood and Seeds appear later, after Ferns have branched off, so Ferns do not share them with Pines.
Therefore, Xylem is the only trait listed that is shared by Ferns and Pines but excluded from Mosses.
An outgroup is a group that branches off from the evolutionary lineage before the other groups (the ingroup).
In Figure 1, the lineage leading to Mosses is the first to diverge from the main stem (bottom left).
This position indicates that Mosses are the outgroup relative to the clade containing Ferns, Pines, and Oaks.
The boxed node represents a branching point (speciation event).
This specific node is the point where the lineage leading to Ferns diverges from the lineage leading to Pines and Oaks.
Therefore, this node represents the most recent common ancestor of the two groups that emerge from it: Ferns and Pines.
(Note: While it is also the ancestor of Oaks, “Ferns and Pines” is the correct pair representing the immediate split at that node).
Cladograms depict relatedness based on common ancestry.
Species \(S\) and species \(T\) share a common ancestor at the node located after trait \(B\) evolved.
Species \(S\) and species \(R\) share a more distant common ancestor at the node before trait \(B\) evolved.
Since \(S\) and \(T\) share a more recent node than \(S\) and \(R\), Species \(S\) is more closely related to species \(T\).
“Derived” in cladistics refers to a trait that evolved later (more recently) relative to ancestral traits.
Looking at the timeline of the cladogram from left (ancestral) to right (recent):
Trait \(B\) appears on the main line before the divergence of species \(S\).
Trait \(C\) appears on the line leading to species \(T\), after the divergence of species \(S\).
Since \(C\) appears later in the evolutionary sequence than \(B\), Characteristic \(C\) is more derived than characteristic \(B\).
Question
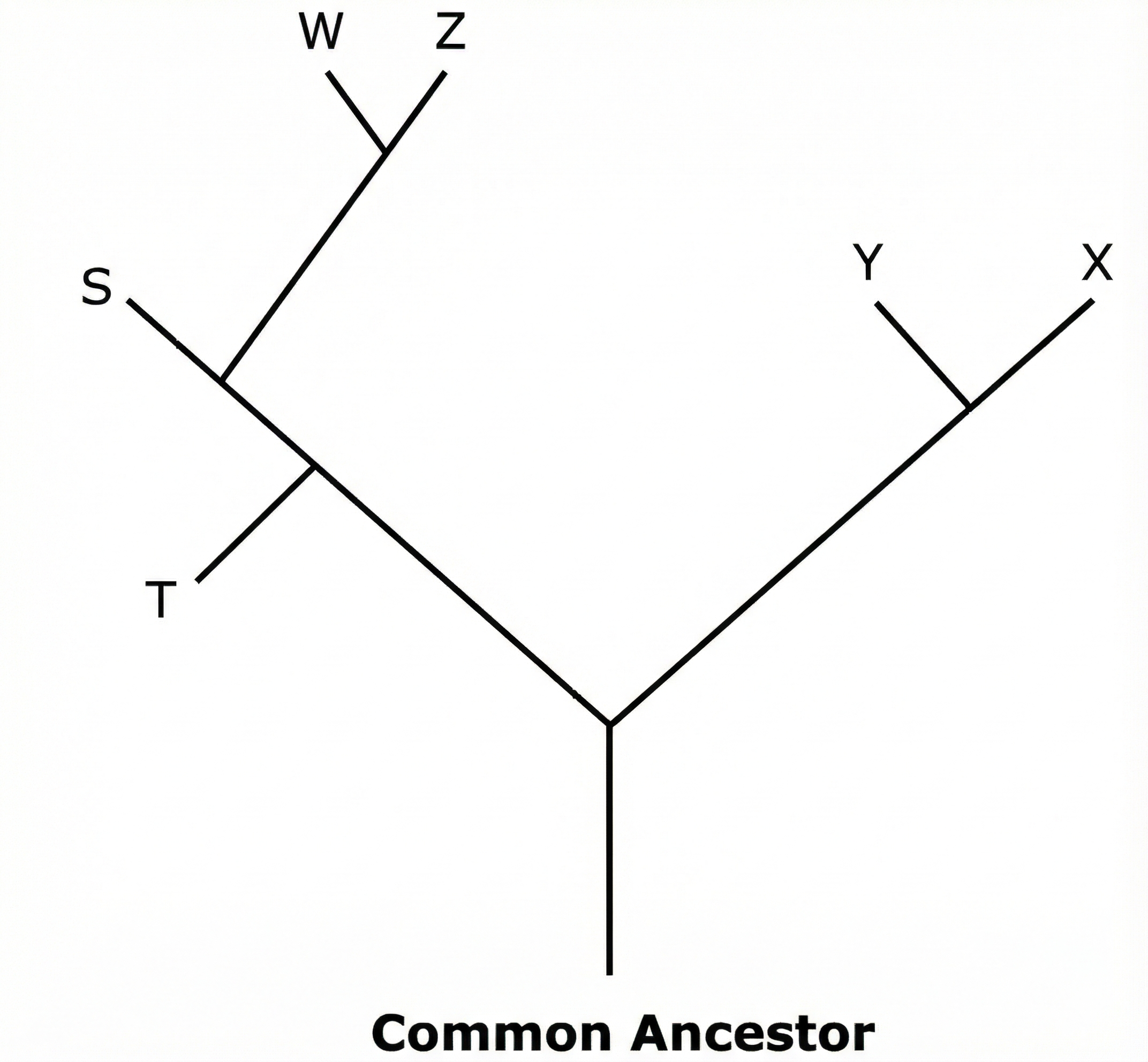
B. organism $T$
C. organism $W$
D. organism $Y$
▶️ Answer/Explanation
In a phylogenetic tree, the vertical or diagonal axis typically represents time.
The base of the tree represents the common ancestor, which is the furthest point in the past.
Organisms that branch off closer to the root (the bottom) appeared earlier in evolutionary history.
Looking at the diagram, the branch for organism $T$ is the first to diverge from the main lineage.
Because it splits off at the lowest point relative to the others, its fossil would be the oldest.
Organisms like $W$, $Z$, $X$, and $Y$ appear at higher nodes, indicating they evolved more recently.
Question
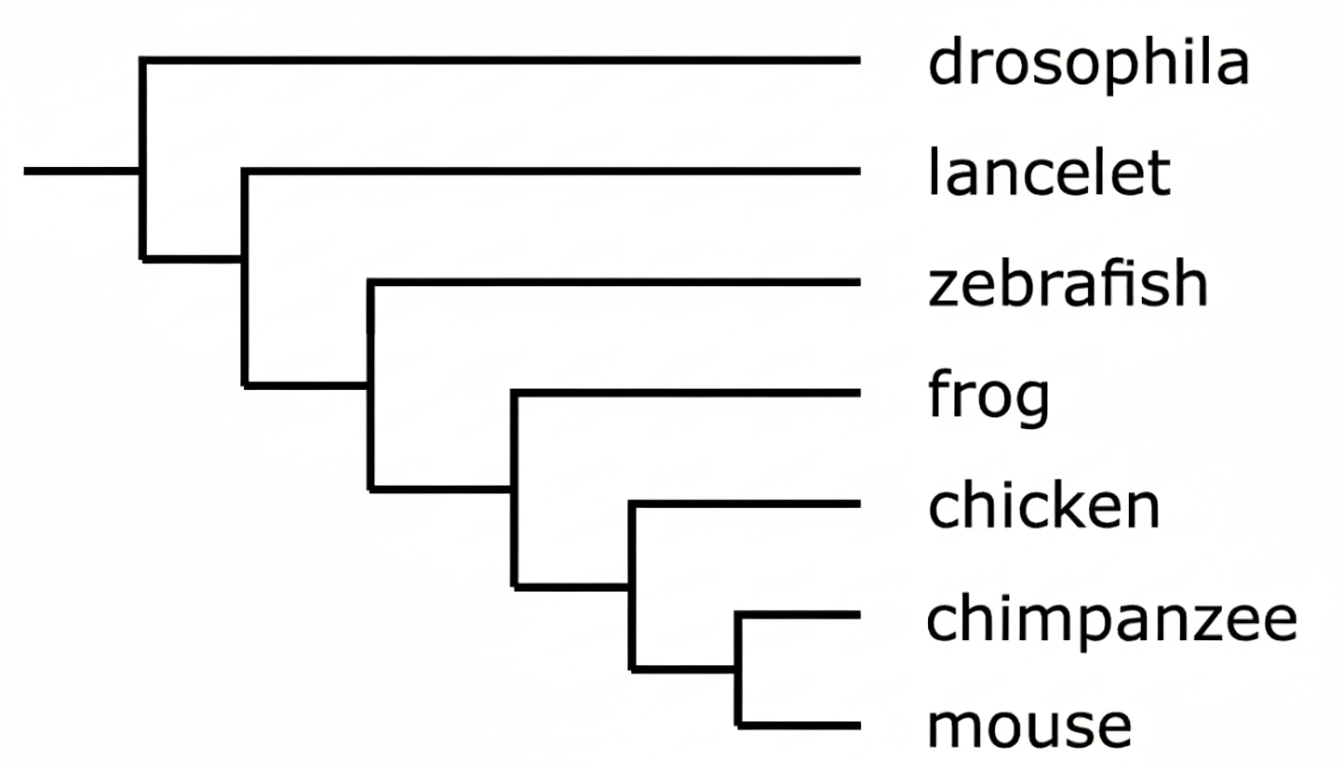
▶️ Answer/Explanation
The correct answer is C.
In a phylogenetic tree, the most recent common ancestor is represented by the node (branching point) closest to the tips.
The node connecting the chicken and the mouse is further to the right than the nodes for other pairs listed.
Specifically, the chimpanzee and mouse share the most recent ancestor overall.
Among the given choices, the chicken and mouse share a more recent ancestor than frogs/chickens or drosophila/lancelets.
This indicates a more recent evolutionary divergence and closer genetic relationship.
Therefore, option C is the most accurate choice based on the branching structure provided.
