Pre AP Biology -GEN 3.4 Mutations- MCQ Exam Style Questions -New Syllabus 2025-2026
Pre AP Biology -GEN 3.4 Mutations- MCQ Exam Style Questions – New Syllabus 2025-2026
Pre AP Biology -GEN 3.4 Mutations- MCQ Exam Style Questions – Pre AP Biology – per latest Pre AP Biology Syllabus.
Question
b. The diseases they cause progress very rapidly.
c. They have an amino acid sequence different from the normal protein.
d. They reproduce by misfolding normal proteins.
▶️ Answer/Explanation
The correct option is d.
Prions are infectious pathogens composed entirely of protein material.
They do not contain nucleic acids like DNA or RNA for replication.
Instead, they “reproduce” by inducing existing normal proteins (\(PrP^C\)) to misfold.
This conformational change converts the normal protein into the infectious form (\(PrP^{Sc}\)).
The amino acid sequence remains identical; only the three-dimensional shape changes.
This process triggers a chain reaction, leading to protein aggregation and brain damage.
Question
b. when exon shuffling occurs
c. when the genes are expressed by transcription and translation
d. when different mutations occur in each protein-coding sequence
▶️ Answer/Explanation
The correct answer is d.
A multigene family starts with a gene duplication event, creating two identical copies.
Evolutionary divergence only begins when independent mutations accumulate in the sequences.
These mutations alter the amino acid sequence, leading to distinct protein functions.
Duplication (option a) provides the raw material but does not itself create functional variety.
Transcription and translation (option c) merely express the existing genetic code.
Therefore, mutational divergence is the specific point where functions start to differ.
Question
▶️ Answer/Explanation
The correct answer is b. a longer mature mRNA.
An insertion adds a nucleotide base to the DNA sequence, which is then transcribed into the pre-mRNA.
Exons are the coding regions that are retained and joined together during the process of RNA splicing.
Because the mutation occurs within an exon, the extra base remains in the final transcript.
Spliceosomes recognize specific sequences at intron-exon boundaries, which are usually unaffected by internal exon insertions.
Translation initiation typically depends on the $5’$ cap and start codon, which remain functional.
A single base insertion shifts the reading frame, making a silent mutation ($d$) virtually impossible.
Question
b. A base that is incorporated into the DNA helix instead of $A$, $T$, $G$, or $C$
c. A base that has an unusual number of hydrogen bonds
d. A base that is found in RNA but not DNA
▶️ Answer/Explanation
The correct answer is b.
Base analogues are chemical compounds that structurally resemble the standard nitrogenous bases ($A$, $T$, $G$, $C$).
Due to this similarity, DNA polymerase may mistakenly incorporate them into the DNA strand during replication.
An example is $5$-bromouracil ($5$-$\text{BU}$), which acts as an analogue to thymine ($T$).
Once incorporated, these analogues can cause mutations by mispairing with non-complementary bases.
Unlike damaged bases (option a) or RNA-specific bases like uracil (option d), analogues are external mutagens.
They shift between tautomeric forms more readily, leading to point mutations in the genetic code.
Question
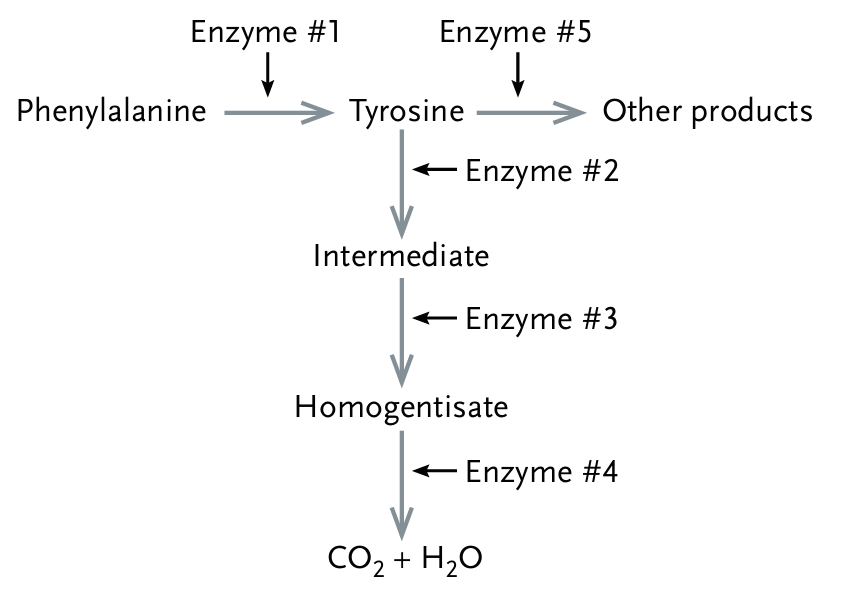
b. A mutation for enzyme $\#2$ prevents tyrosine from being synthesized.
c. A mutation at enzyme $\#3$ prevents homogentisate from being synthesized.
d. A mutation for enzyme $\#4$ could hide a mutation in enzyme $\#1$.
▶️ Answer/Explanation
The correct option is c.
Enzyme $\#3$ catalyzes the conversion of the Intermediate into Homogentisate.
If a mutation destroys enzyme $\#3$, the pathway is blocked at that specific step.
Consequently, Homogentisate cannot be produced, leading to a deficiency of that product.
Option a is incorrect because enzyme $\#1$ produces tyrosine; its loss would decrease tyrosine.
Option b is incorrect because tyrosine is a precursor to enzyme $\#2$, not a product of it.
Option d is incorrect because a downstream mutation (enzyme $\#4$) cannot mask the loss of an upstream product (from enzyme $\#1$).
Question
▶️ Answer/Explanation
The correct answer is D.
A point mutation occurs when a single nucleotide base is changed.
Specifically, it involves the substitution of one nitrogenous base for another.
Unlike insertions or deletions, a substitution only affects a single codon.
Options B and C describe chromosomal mutations involving large DNA segments.
Option A refers to frameshift mutations which alter the entire reading frame.
Therefore, the precise definition of a point mutation is the replacement of 1 base.
Question
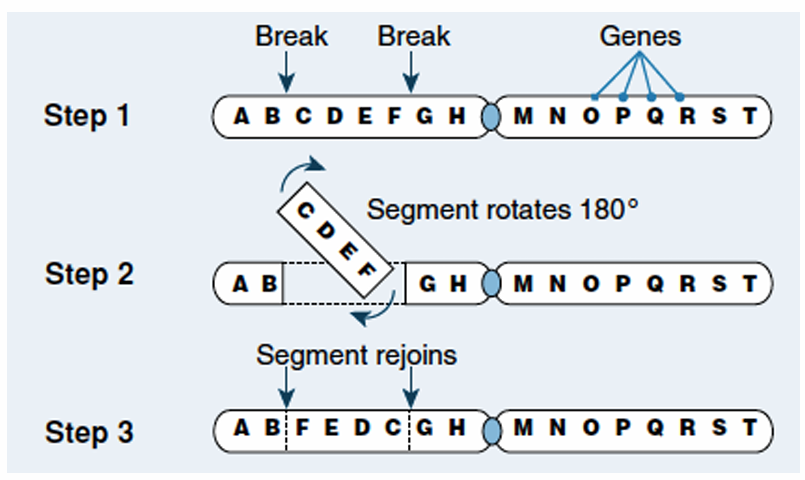
B. translocation.
C. inversion.
D. duplication.
▶️ Answer/Explanation
The correct answer is C. inversion.
In Step $1$, two breaks occur within a single chromosome arm.
The segment containing genes $C$, $D$, $E$, and $F$ is detached.
In Step $2$, this specific segment rotates $180^{\circ}$.
In Step $3$, the segment rejoins the chromosome in reverse order.
The resulting gene sequence changes from $A-B-C-D-E-F-G-H$ to $A-B-F-E-D-C-G-H$.
Because the segment is flipped rather than lost or moved elsewhere, it is defined as an inversion.
Question
▶️ Answer/Explanation
The correct answer is B. Only primary and tertiary structure.
Every protein has a primary structure, which is the unique sequence of amino acids in the polypeptide chain.
Since the protein is described as “highly complex and folded,” it must possess a tertiary structure, which is the overall three-dimensional shape.
The prompt explicitly states there are no alpha ($\alpha$) helices or beta ($\beta$) sheets, which means there is no secondary structure present.
Because it consists of a single polypeptide, it cannot have a quaternary structure, as that requires multiple subunits.
Therefore, the only levels of organization present are the primary and tertiary structures.
Question
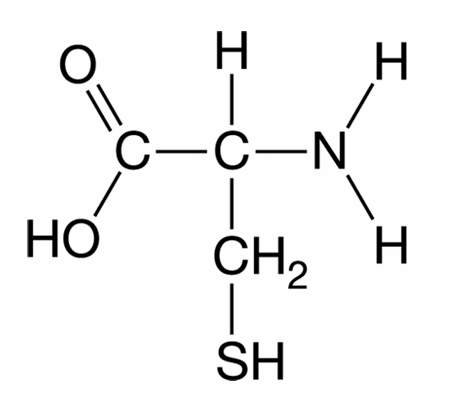
B. Tertiary structure.
C. Quaternary structure.
D. Either tertiary or quaternary structure depending upon if the cysteine—cysteine $R$-side chain bonds are between amino acids in the same polypeptide or two different polypeptides.
▶️ Answer/Explanation
The correct option is D.
Cysteine contains a sulfhydryl ($-SH$) group in its $R$-group side chain.
Two cysteine side chains can oxidize to form a covalent disulfide bridge ($-S-S-$).
If this bond forms within a single polypeptide chain, it stabilizes the tertiary structure.
If the bond forms between different polypeptide subunits, it stabilizes the quaternary structure.
Secondary structure is primarily stabilized by hydrogen bonds between the backbone, not $R$-groups.
Therefore, the classification depends entirely on whether the bond is intra-chain or inter-chain.
Question
B. The presence of two or more polypeptides connected by $R$-side chain bonding.
C. Structures such as alpha-helices that are made by backbone hydrogen bonding.
D. The order or sequence of amino acids in a polypeptide chain.
▶️ Answer/Explanation
The correct answer is C.
Secondary structure refers to local folding patterns within a polypeptide.
It is stabilized specifically by hydrogen bonds between atoms of the peptide backbone.
Common examples include the $\alpha$-helix and the $\beta$-pleated sheet.
Option A describes tertiary structure involving $R$-group interactions.
Option B describes quaternary structure involving multiple subunits.
Option D describes the primary structure or amino acid sequence.
